-
 Caffeic Acid and Biopesticides Interactions for the Control of Storage Beetles
Caffeic Acid and Biopesticides Interactions for the Control of Storage Beetles -
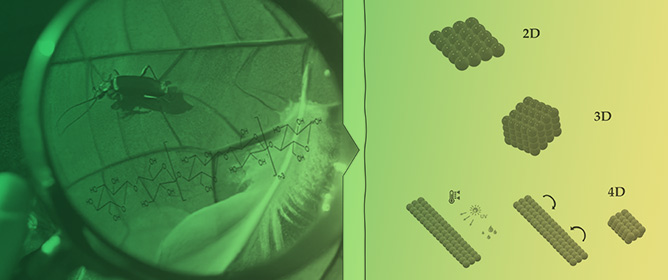 Nature-Inspired Cellulose-Based Active Materials: From 2D to 4D
Nature-Inspired Cellulose-Based Active Materials: From 2D to 4D -
 How Far Are We from Research That Is Independent of the Use of Animal Models? A Comparative Analysis between Animal and 3D/On-a-Chip Models for the Study of Respiratory Diseases
How Far Are We from Research That Is Independent of the Use of Animal Models? A Comparative Analysis between Animal and 3D/On-a-Chip Models for the Study of Respiratory Diseases -
 Crossiella, a Rare Actinomycetota Genus, Abundant in the Environment
Crossiella, a Rare Actinomycetota Genus, Abundant in the Environment
Journal Description
Applied Biosciences
Applied Biosciences
is an international, peer-reviewed, open access journal on all aspects of applied biosciences published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: first decisions in 16 days; acceptance to publication in 5.8 days (median values for MDPI journals in the first half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- Applied Biosciences is a companion journal of Applied Sciences.
Latest Articles
Comparative Analysis of Bioactive Phenolic Compounds and Fatty Acids in Seeds and Seedlings of Canadian Alfalfa, Sainfoin, and Fenugreek
Appl. Biosci. 2023, 2(3), 477-492; https://doi.org/10.3390/applbiosci2030030 (registering DOI) - 01 Sep 2023
Abstract
►
Show Figures
The interest in under-utilized crops as a functional food for animals and humans has been increasing recently with advancing research and the need for crop improvement. Canadian forage crops including alfalfa (Medicago sativa L.) and fenugreek (Trigonella foenum-graecum L.) are
[...] Read more.
The interest in under-utilized crops as a functional food for animals and humans has been increasing recently with advancing research and the need for crop improvement. Canadian forage crops including alfalfa (Medicago sativa L.) and fenugreek (Trigonella foenum-graecum L.) are marketed in various forms due to their traditionally known health benefits. Sainfoin (Onobrychis viciifolia Scop.) is another forage crop with potential health benefits containing beneficial nutraceuticals. In this study, we assessed selected bioactive phenolic compounds and fatty acids in seeds and seedlings of Canadian-grown alfalfa, sainfoin, and fenugreek. Various phenolic compounds were detected in all three forage crop seeds and seedlings. In general, Sainfoin seeds were high in phenolic compounds relative to that of alfalfa and fenugreek. Chlorogenic acid, epigallo catechin, and gallic acid were at high concentrations at 56.6, 86.8, and 64.7 µg.g−1, respectively, compared to other phenolic compounds in sainfoin seeds. The fatty acids content (%) was significantly affected by the seedling stage and crop type. Some of the bioactive compounds present in seeds were not detected in seedling stages. The comparative bioactive phenolic compounds and fatty acid assessments of these forage legumes could potentially be used as biomarkers for the selection and development of favorable cultivars for animal and human nutrition. In addition, these crops could be used for isolating these bioactive compounds, and thus increasing their agri-food value.
Full article
Open AccessFeature PaperReview
Deregulations of RNA Pol II Subunits in Cancer
Appl. Biosci. 2023, 2(3), 459-476; https://doi.org/10.3390/applbiosci2030029 - 24 Aug 2023
Abstract
►▼
Show Figures
Deregulated transcription is a well-known characteristic of cancer cells, with differentially expressed genes being a common feature of several cancers. Often, deregulated transcription is a consequence of alterations in transcription factors (TFs), which play a crucial role in gene expression and can act
[...] Read more.
Deregulated transcription is a well-known characteristic of cancer cells, with differentially expressed genes being a common feature of several cancers. Often, deregulated transcription is a consequence of alterations in transcription factors (TFs), which play a crucial role in gene expression and can act as tumour suppressors or proto-oncogenes. In eukaryotic organisms, transcription is carried out by three distinct RNA polymerase complexes: Pol I, Pol II, and Pol III. Pol II, specifically, is responsible for transcribing messenger RNA (mRNA), the protein coding part of the genome, as well as long non-coding RNAs (lncRNAs). While there is considerable research on the impact of specific deregulated transcription factors in cancer development, there is a lack of studies focusing on defects within the RNA polymerase complexes and their subunits. This review aims to shed light in particular on the Pol II complex and highlight the deregulation of its subunits that have a significant impact on tumour development, prognosis, and survival. By providing a comprehensive overview of our current understanding of Pol II subunits in cancer, this review emphasizes the importance of further research in this area. It suggests that exploring these subunits’ deregulations could lead to the identification of valuable biomarkers and potential therapeutic targets, making it a topic of collective interest.
Full article

Figure 1
Open AccessArticle
Sequencing, Fast and Slow: Profiling Microbiomes in Human Samples with Nanopore Sequencing
Appl. Biosci. 2023, 2(3), 437-458; https://doi.org/10.3390/applbiosci2030028 - 17 Aug 2023
Abstract
Rapid and accurate pathogen identification is crucial in effectively combating infectious diseases. However, the current diagnostic tools for bacterial infections predominantly rely on century-old culture-based methods. Furthermore, recent research highlights the significance of host–microbe interactions within the host microbiota in influencing the outcome
[...] Read more.
Rapid and accurate pathogen identification is crucial in effectively combating infectious diseases. However, the current diagnostic tools for bacterial infections predominantly rely on century-old culture-based methods. Furthermore, recent research highlights the significance of host–microbe interactions within the host microbiota in influencing the outcome of infection episodes. As our understanding of science and medicine advances, there is a pressing need for innovative diagnostic methods that can identify pathogens and also rapidly and accurately profile the microbiome landscape in human samples. In clinical settings, such diagnostic tools will become a powerful predictive instrument in directing the diagnosis and prognosis of infectious diseases by providing comprehensive insights into the patient’s microbiota. Here, we explore the potential of long-read sequencing in profiling the microbiome landscape from various human samples in terms of speed and accuracy. Using nanopore sequencers, we generate native DNA sequences from saliva and stool samples rapidly, from which each long-read is basecalled in real-time to provide downstream analyses such as taxonomic classification and antimicrobial resistance through the built-in software (<12 h). Subsequently, we utilize the nanopore sequence data for in-depth analysis of each microbial species in terms of host–microbe interaction types and deep learning-based classification of unidentified reads. We find that the nanopore sequence data encompass complex information regarding the microbiome composition of the host and its microbial communities, and also shed light on the unexplored human mobilome including bacteriophages. In this study, we use two different systems of long-read sequencing to give insights into human microbiome samples in the ‘slow’ and ‘fast’ modes, which raises additional inquiries regarding the precision of this novel technology and the feasibility of extracting native DNA sequences from other human microbiomes.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures
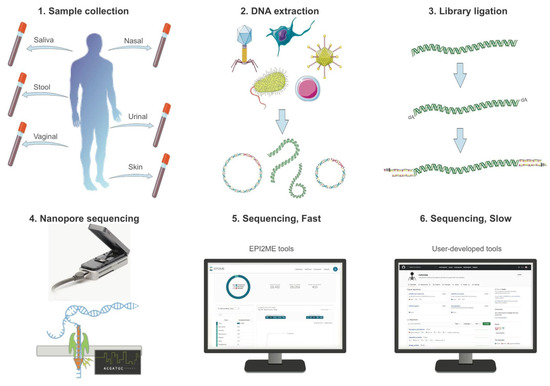
Figure 1
Open AccessArticle
Analysis of Adolescents’ Head to Shoulder Region during Tablet Use from Sagittal and Frontal RGB Images
by
and
Appl. Biosci. 2023, 2(3), 421-436; https://doi.org/10.3390/applbiosci2030027 - 04 Aug 2023
Abstract
►▼
Show Figures
As schools go digital, the use of tablet computers is increasing. Concerns are raised that the extensive use of tablets and the associated bent-over posture may negatively affect the individual’s health. In order to analyse the possible effects of prolonged tablet use on
[...] Read more.
As schools go digital, the use of tablet computers is increasing. Concerns are raised that the extensive use of tablets and the associated bent-over posture may negatively affect the individual’s health. In order to analyse the possible effects of prolonged tablet use on physical health, a detailed analysis of the posture during tablet use is needed so that appropriate preventive measures can be taken to prevent degenerative changes. Therefore, the aim of this study was to measure and report the posture of 56 students while working with a tablet computer and compare it with an upright posture. Sagittal and frontal images were used for measurements of the subjects’ postures while seated, using the tablet, and in a neutral sitting position looking straight ahead. The body position during tablet use was recorded in two different user configurations: tablet flat on the table and tablet in individual freely chosen user configuration. After appropriate annotation of the data, the following parameters were evaluated in different planes. The craniovertebral angle (CVA), head tilt angle (HTA), and forward shoulder angle (FSA) are measurements that describe the extent to which the head bends forward and downward and how the shoulders are aligned in the sagittal plane. On the other hand, the head shoulder angle (HSA), lateral head tilt angle (LHTA), and trunk flexion angle (TFA) are angles measured in the frontal plane, which indicate the degree of head tilt and trunk bending to the right or left side. The measurement results clearly showed that the use of a tablet had a pronounced effect on the positions and rotations of the participants’ head, neck, and shoulders. This was evident through strong deviations observed in the angles measured between the sitting straight posture and the postures while using the tablet. For example, depending on the body posture class, the mean CVA values were 45.76° for straight sitting posture, 28.25° for holding the tablet individually posture, and 26.04° for the posture adopted while using a tablet placed flat on the table.
Full article

Figure 1
Open AccessArticle
Effect of Doxapram, a K2p Channel Blocker, and pH on Heart Rate: Larval Drosophila Model
Appl. Biosci. 2023, 2(3), 406-420; https://doi.org/10.3390/applbiosci2030026 - 03 Aug 2023
Abstract
►▼
Show Figures
Two-P-domain K+ (K2p) channels are responsible for maintaining the resting membrane potential. K2p channels have varied expression in healthy tissue, but they also change in cancerous or diseased states. The correlation and causation as regards the alteration of K2p channel expression are
[...] Read more.
Two-P-domain K+ (K2p) channels are responsible for maintaining the resting membrane potential. K2p channels have varied expression in healthy tissue, but they also change in cancerous or diseased states. The correlation and causation as regards the alteration of K2p channel expression are still being investigated. The compound doxapram seems to block K2p channels and depolarize cells. Using Drosophila, the increased expression of the ORK1 K2p channel in cardiac and skeletal muscle was investigated. The heart rate in larval Drosophila is very sensitive to pH, and since doxapram blocks a subset of the K2p channels that are known to be acid-sensitive, it was postulated that doxapram would affect heart rate. A pH change from 7.1 to 6.5 increased the rate, while that from 7.1 to 7.5 decreased the rate. An amount of 0.1 mM of doxapram had no effect, but 0.5 of mM depressed Drosophila heart rates within five minutes. Exposure to 5 mM of doxapram immediately decreased the rate. Lipopolysaccharides (LPSs) from Gram-negative bacteria acutely increased the rate. LPSs activate K2p channels in the skeletal muscle of larvae and are blocked by doxapram. LPSs slightly reduce depression in the rate induced by doxapram. The overexpression of K2p channels in the heart and skeletal muscle depressed the heart rate and heightened pH sensitivity. At larval neuromuscular junctions, the overexpression in skeletal muscle increases the frequency of spontaneous quantal events and produces a more negative resting membrane potential.
Full article

Figure 1
Open AccessArticle
Automation of Cluster Extraction in Fundus Autofluorescence Images of Geographic Atrophy
by
and
Appl. Biosci. 2023, 2(3), 384-405; https://doi.org/10.3390/applbiosci2030025 - 21 Jul 2023
Abstract
►▼
Show Figures
The build-up of lipofuscin—an age-associated biomarker referred to as hyperfluorescence—is considered a precursor in the progression of geographic atrophy (GA). Prior studies have attempted to classify hyperfluorescent regions to explain varying rates of GA progression. In this study, digital image processing and unsupervised
[...] Read more.
The build-up of lipofuscin—an age-associated biomarker referred to as hyperfluorescence—is considered a precursor in the progression of geographic atrophy (GA). Prior studies have attempted to classify hyperfluorescent regions to explain varying rates of GA progression. In this study, digital image processing and unsupervised learning were used to (1) completely automate the extraction of hyperfluorescent regions from images, and (2) evaluate prospective patterns and groupings of hyperfluorescent areas associated with varying levels of GA progression. Patterns were determined by clustering methods, such as k-Means, and performance was evaluated using metrics such as the Silhouette Coefficient (SC), the Davies–Bouldin Index (DBI), and the Calinski–Harabasz Index (CHI). Automated extraction of hyperfluorescent regions was carried out using pseudocoloring techniques. The approach revealed three distinct types of hyperfluorescence based on color intensity changes: early-stage hyperfluorescence, intermediate-stage hyperfluorescence, and late-stage hyperfluorescence, with the early and late stages having three additional subclassifications that could explain varying levels of GA progression. The performance metrics for early-stage hyperfluorescence were SC = 0.597, DBI = 0.915, and CHI = 186.989. For late-stage hyperfluorescence, SC = 0.593, DBI = 1.013, and CHI = 217.325. No meaningful subclusters were identified for the intermediate-stage hyperfluorescence, possibly because it is a transitional phase of hyperfluorescence progression.
Full article
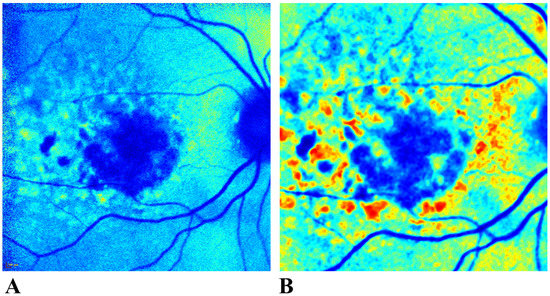
Figure 1
Open AccessCommunication
Review of Partial Hybrids between Herbaceous Medicago sativa and Woody Medicago arborea and Their Potential Role in Alfalfa Improvement
by
and
Appl. Biosci. 2023, 2(3), 373-383; https://doi.org/10.3390/applbiosci2030024 - 13 Jul 2023
Abstract
Medicago sativa (2n = 4x = 32) and M. arborea (2n = 4x = 32) were thought to be reproductively isolated until hybrids (Alborea) were produced by sexual reproduction for the first time in 2003 in Wisconsin. The hybrids were asymmetric, at or
[...] Read more.
Medicago sativa (2n = 4x = 32) and M. arborea (2n = 4x = 32) were thought to be reproductively isolated until hybrids (Alborea) were produced by sexual reproduction for the first time in 2003 in Wisconsin. The hybrids were asymmetric, at or near 2n = 4x = 32, and with a predominance of the alfalfa genome. Only M. sativa seed parents with reproductive abnormalities, including unreduced eggs, have produced hybrids; where M. arborea has been used as the seed parent, no hybrids have resulted. Pedigree selection within derivatives of the two original M. sativa seed parents (MB and M8) has been successful in increasing the frequency of hybrids produced. While Alborea individuals more closely resemble M. sativa, a number of M. arborea-specific traits have been observed across different hybrid individuals. These include single-coil flat pods, large seeds, yellow flowers, indeterminate growth, a minimal crown, lodging, frost resistance, and anthracnose resistance. These M. arborea traits have the potential to restructure alfalfa to increase its versatility and utilisation. There is emerging evidence from North and South America and Australia that some Alborea selections have the capacity to complement adapted alfalfa cultivars for yield. Work is continuing to introgress M. arborea traits of value into alfalfa.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures
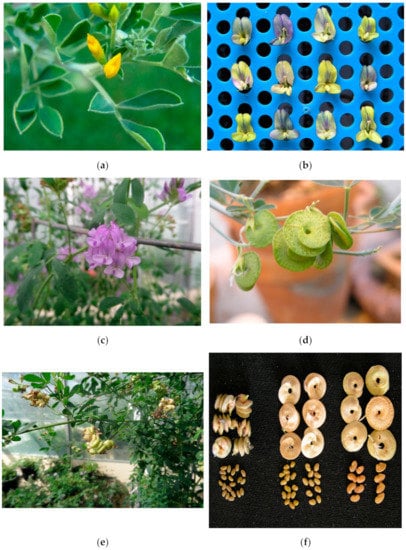

Figure 1
Open AccessReview
Wine Grapes Ripening: A Review on Climate Effect and Analytical Approach to Increase Wine Quality
Appl. Biosci. 2023, 2(3), 347-372; https://doi.org/10.3390/applbiosci2030023 - 10 Jul 2023
Abstract
Red wine grapes have an important impact on the economy of many regions, both for wine quality and for their richness in phenolic compounds, which have many health benefits. Climate has been changing substantially in the last years, which affects greatly grape polyphenolic
[...] Read more.
Red wine grapes have an important impact on the economy of many regions, both for wine quality and for their richness in phenolic compounds, which have many health benefits. Climate has been changing substantially in the last years, which affects greatly grape polyphenolic composition and wine quality. In this review, we will unveil the importance of climate in grape development, both physically and chemically, the different methodologies used to evaluate grape quality, the interesting new approaches using NIR spectroscopy, and the functional properties of grapes and red wine, due to their high phenolic content. Climate has an impact in the development of phenolic compounds in grapes, namely in the anthocyanins biosynthesis. The phenolic chemical composition changes during maturation, therefore, it is essential to keep on track the accumulation of these key compounds. This information is crucial to help producers choose the best harvest date since specific compounds like polyphenols are responsible for the color, taste, and mouthfeel of wines, which directly affects wine quality. The usage of different methodologies to assess quality parameters in grapes and wine, can be used to provide essential information to create the chemical profile of each variety to develop calibration methods. NIR spectroscopy seems to be a reliable method to be used in vineyards during grape maturation to provide real time information on quality parameters to producers since many reliable calibration models have been developed over time.
Full article
(This article belongs to the Special Issue Plant Natural Compounds: From Discovery to Application)
►▼
Show Figures
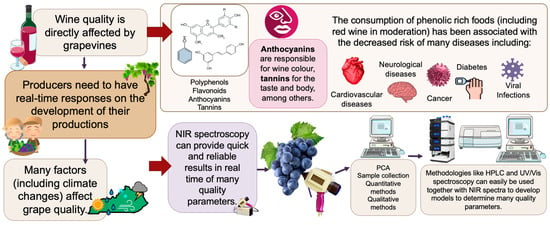
Figure 1
Open AccessArticle
Exploring the Synergistic Impacts of Cover Crops and Fertilization on Soil Microbial Metabolic Diversity in Dryland Soybean Production Systems Using Biolog EcoPlates
by
, , , and
Appl. Biosci. 2023, 2(3), 328-346; https://doi.org/10.3390/applbiosci2030022 - 06 Jul 2023
Abstract
The metabolic diversity of soil microbiota embodies diverse functional capabilities that support ecosystem resilience, driving essential biogeochemical processes and facilitating the optimization of sustainable agricultural systems. Integrating cover crops into agricultural systems cultivates a diverse array of metabolic activities among soil microbes, synergistically
[...] Read more.
The metabolic diversity of soil microbiota embodies diverse functional capabilities that support ecosystem resilience, driving essential biogeochemical processes and facilitating the optimization of sustainable agricultural systems. Integrating cover crops into agricultural systems cultivates a diverse array of metabolic activities among soil microbes, synergistically enhancing ecosystem services and bolstering soil health for sustainable and productive farming practices. In an effort to gain deeper insights and expand our knowledge, we conducted a study examining the effects of cover crops and fertilizer sources, thereby shedding light on their combined impacts on the metabolic activity dynamics of soil microbial communities. In this investigation, we employed a split-plot design with two factors: (a) cover crop with three solo cover crop species—Cereal rye (Secale cereale), wheat (Triticum aestivum), hairy vetch (Vicia villosa), and one mixture of mustard (Brassica rapa) and cereal rye (Secale cereale) (CC-mix), (b) Fertilizer source includes poultry litter, chemical fertilizer, and no-fertilizer treatments. We assessed the metabolic potential of soil microbiota by using carbon substrates utilizing Biolog EcoPlates. The findings revealed that the plots with CC-mix treatment exhibited greater metabolic diversity compared to the other treatments, while among the fertilizer sources, poultry litter demonstrated higher metabolic activity. Furthermore, both treatment factors predominantly metabolized carbohydrates and polymers compared to other carbon substrate categories. The principal component analysis accounted for 46.4% of the variance, collectively represented by PC1 and PC2, emphasizing the substantial contributions of carbohydrates, amino acids, and carboxylic acids to the observed metabolic diversity. Canonical correspondence analysis revealed that pH had positively correlated with microbial functional diversity, whereas total carbon (TC), total nitrogen (TN), and water-stable aggregates (WSA) showed a negative correlation. In conclusion, cover cropping and type of fertilizer source had a notable impact on soil microbial functional diversity, with the cover crop mixture exhibiting a more pronounced influence than the individual cover crop treatments.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures
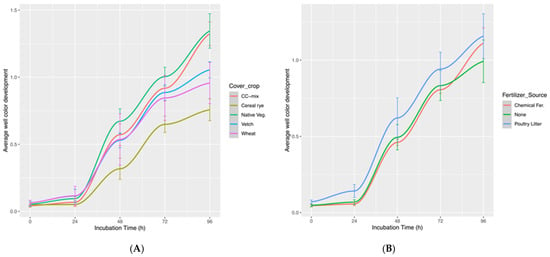
Figure 1
Open AccessArticle
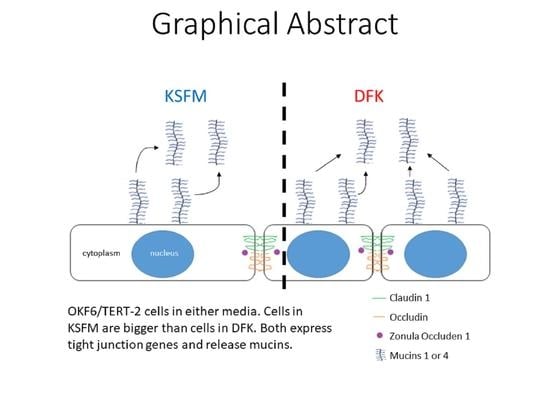
Comparison of Culture Media for In Vitro Expansion of Oral Epithelial Keratinocytes
Appl. Biosci. 2023, 2(2), 308-327; https://doi.org/10.3390/applbiosci2020021 - 16 Jun 2023
Abstract
►▼
Show Figures
Background: Expansion of OKF6/TERT-2 oral epithelial cells in vitro is important for studying the molecular biology of disease and pathology affecting the oral cavity. Keratinocyte serum-free medium (KSFM) is the medium of choice for this cell line. This study compares three media for
[...] Read more.
Background: Expansion of OKF6/TERT-2 oral epithelial cells in vitro is important for studying the molecular biology of disease and pathology affecting the oral cavity. Keratinocyte serum-free medium (KSFM) is the medium of choice for this cell line. This study compares three media for OKF6/TERT-2 cultures: KSFM, Dulbecco’s Modified Eagle Medium/Nutrient Mixture of Hams F-12 (DMEM/F12), and a composite medium comprised of DMEM/F-12 and KSFM (1:1 v/v), referred to as DFK. The toxicological effects of electronic cigarette liquids (e-liquids) on OKF6/TERT-2 cells cultured in these media were also compared. Methods: Cells were cultured in KSFM, DMEM/F12, or DFK, and cellular morphology, growth, wound healing and the gene expression of mucins and tight junctions were evaluated. Additionally, cytotoxicity was determined after e-liquid exposures. Results: Switching from KSFM to DMEM/F12 or DFK 24 h post-seeding leads to typical cellular morphologies, and these cultures reach confluency faster than those in KSFM. Wound-healing recovery occurred fastest in DFK. Except for claudin-1, there is no difference in expression of the other genes tested. Additionally, e-liquid cytotoxicity appears to be amplified in DFK cultures. Conclusions: DMEM/F12 and DFK are alternative media for OKF6/TERT-2 cell culture to study the molecular biology of disease and pathology, provided cells are initially seeded in KSFM.
Full article
Graphical abstract
Open AccessArticle
An Event-Driven Architecture for Genomics-Based Diagnostic Data Processing
by
, , , , , , , and
Appl. Biosci. 2023, 2(2), 292-307; https://doi.org/10.3390/applbiosci2020020 - 02 Jun 2023
Abstract
►▼
Show Figures
Genomics-based diagnostic data (GBDD) are becoming increasingly important for laboratory diagnostics. Due to the large quantity of data and their heterogeneity, GBDD poses a big data challenge. Current analysis tools for GBDD are primarily designed for research and do not meet the requirements
[...] Read more.
Genomics-based diagnostic data (GBDD) are becoming increasingly important for laboratory diagnostics. Due to the large quantity of data and their heterogeneity, GBDD poses a big data challenge. Current analysis tools for GBDD are primarily designed for research and do not meet the requirements of laboratory diagnostics for automation, reliability, transparency, reproducibility, robustness, and accessibility. This makes it difficult for laboratories to use these tools in tests that need to be validated according to regulatory frameworks and to execute tests in a time- and cost-efficient manner. In order to better address these requirements, we propose an event-driven workflow-based architecture as the basis for a processing platform that is highly scalable using container technologies and microservices. A prototype implementation of this approach, called GenomicInsights, has been developed and evaluated to demonstrate its feasibility and suitability for laboratory diagnostics.
Full article
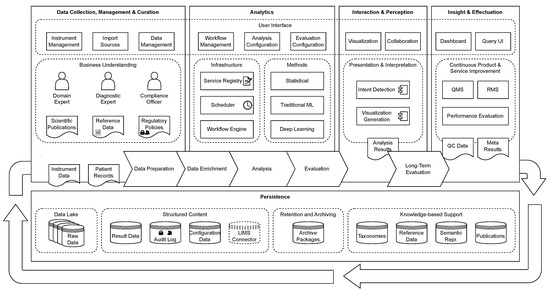
Figure 1
Open AccessArticle
Coupled Biogas and Fiber Production from Agricultural Residues and Energy Crops with Steam Explosion Treatment
by
, , , , , , and
Appl. Biosci. 2023, 2(2), 278-291; https://doi.org/10.3390/applbiosci2020019 - 01 Jun 2023
Abstract
The global demand for packaging materials and energy is constantly increasing, requiring the exploration of new concepts. In this work, we presented a bioeconomic concept that uses steam explosion and phase separation to simultaneously generate fibers for the packaging industry and biogas substrate
[...] Read more.
The global demand for packaging materials and energy is constantly increasing, requiring the exploration of new concepts. In this work, we presented a bioeconomic concept that uses steam explosion and phase separation to simultaneously generate fibers for the packaging industry and biogas substrate for the energy sector. The concept focused on fiber-rich residues and fiber-rich ecological energy crops from agriculture. Feasibility of the concept in the laboratory using feedstocks, including Sylvatic silphia silage, Nettle silage, Miscanthus, Apple pomace, Alfalfa stalks, and Flax shives was confirmed. Our results showed that we were able to separate up to 26.2% of the methane potential while always extracting a smaller percentage of up to 17.3% of organic dry matter (ODM). Specific methane yields of 297–486 LCH4 kgODM−1 in the liquid and 100–286 LCH4 kgODM−1 in the solid phase were obtained. The solid phases had high water absorption capacities of 216–504% due to the steam explosion, while the particle size was not significantly affected. The concept showed high potential, especially for undried feedstock.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures

Figure 1
Open AccessArticle
Computational and Experimental Investigation of the Combined Effect of Various 3D Scaffolds and Bioreactor Stimulation on Human Cells’ Feedback
by
, , , , , , and
Appl. Biosci. 2023, 2(2), 249-277; https://doi.org/10.3390/applbiosci2020018 - 01 Jun 2023
Abstract
Computational methods were combined with an experimental setup in order to investigate the response of human umbilical cord stem cells to 3D electrospun and printed scaffolds, when dynamically stimulated in a bioreactor. Key parameters associated to bioreactor working conditions were computationally investigated using
[...] Read more.
Computational methods were combined with an experimental setup in order to investigate the response of human umbilical cord stem cells to 3D electrospun and printed scaffolds, when dynamically stimulated in a bioreactor. Key parameters associated to bioreactor working conditions were computationally investigated using Comsol software to use the output for the planned experimental setup. Based on the theoretical observations, the influence of the inlet velocity, cell number, and exposure time in the bioreactor were analyzed and the in vitro parameters were adjusted accordingly. MSCs were seeded in different numbers in the 3D porous scaffolds and stimulated in the bioreactor (0.5 and 2 h duration, 3 and 6 mm/s inlet velocity). Polycaprolactone 3D electrospun, and polyurethane and polylactic acid 3D-printed scaffolds were fabricated and fibronectin-coated. The computational study predicted initial events in the process of cells deposition and attachment. Total protein, osteopontin, and osteocalcin levels in cells deposited in scaffolds were investigated; SEM and confocal imaging confirmed the biomarker analysis. MSCs proliferated well in PCL. Polyurethane enabled extremely rapid proliferation followed by differentiation, while PLA induced a moderate proliferation and parallel mineralization. The scaffolds stiffness has been found as the key enabling parameter decisive for cells feedback.
Full article
(This article belongs to the Special Issue Anatomy and Regenerative Medicine: From Methods to Applications)
►▼
Show Figures
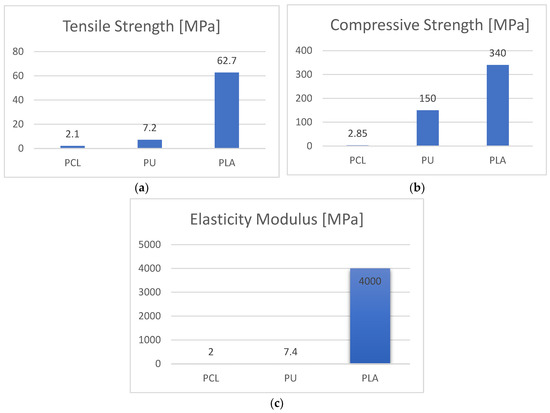
Figure 1
Open AccessArticle
Effect of 2-Aminoethoxydiphenyl Borate (2-APB) on Heart Rate and Relation with Suppressed Calcium-Activated Potassium Channels: Larval Drosophila Model
Appl. Biosci. 2023, 2(2), 236-248; https://doi.org/10.3390/applbiosci2020017 - 23 May 2023
Abstract
►▼
Show Figures
Cardiac contractile cells depend on calcium in order to function. Understanding the regulation of calcium influx, efflux, and release from the sarcoplasmic reticulum is essential. The focus of this investigation is to address how a reduction of functional Ca2+-activated K+
[...] Read more.
Cardiac contractile cells depend on calcium in order to function. Understanding the regulation of calcium influx, efflux, and release from the sarcoplasmic reticulum is essential. The focus of this investigation is to address how a reduction of functional Ca2+-activated K+ (KCa) channels, via a mutational line, might impact the heart rate in larva when the SER is also modulated through Ca2+ loading and stimulation. The larval heart tube is exposed in situ and flushed with saline. With a known saline composition, a potential therapeutic pharmacological agent, 2-Aminoethyl diphenylborinate (2-APB), was examined for its effect on heart rate, as well as to determine the contribution from KCa channels. In this study, it was determined that mutation in the K(Ca) channel (i.e., Slo) showed a different trend than the wild-type CS strain. Exposure to high concentrations of 50 µM 2-APB decreased heart rate in the Slo strain and increased it in the wild-type CS strain. Serotonin increased heart rate in both thapsigargin- and 2-APB-treated larvae, with no significant difference between the strains.
Full article
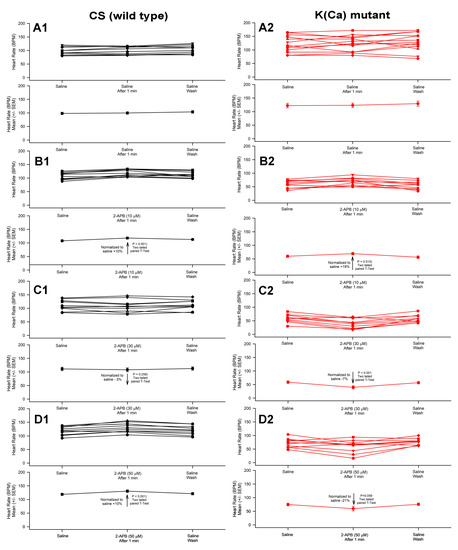
Figure 1
Open AccessArticle
Seed Morpho-Anatomy and Germination Enhancement of the Australian Native Species Lomandra longifolia Labill. and L. hystrix (R.Br.) L.R. Fraser & Vickery
by
, , , and
Appl. Biosci. 2023, 2(2), 222-235; https://doi.org/10.3390/applbiosci2020016 - 16 May 2023
Abstract
Lomandra species are an important understory component of many Australian native ecosystems, contributing to the floristic richness and stabilizing soils. However, a limited understanding of their germination biology currently hinders their efficient use in seed-based restoration and ornamental plant production. The present study
[...] Read more.
Lomandra species are an important understory component of many Australian native ecosystems, contributing to the floristic richness and stabilizing soils. However, a limited understanding of their germination biology currently hinders their efficient use in seed-based restoration and ornamental plant production. The present study investigated Lomandra longifolia and L. hystrix diaspore morpho-anatomy and evaluated different mechanical and/or chemical treatments (nicking, leaching, smoke water and gibberellic acid [GA3]) and under light or dark conditions to enhance germination. Embryos of both species were small and linear with a low embryo to seed ratio (<0.45). Germination rates of both species were significantly hastened by leaching seeds in running water for 36 h as compared to a non-leached seed. The results suggest that pre-treating both Lomandra species by leaching could maximize the effectiveness of seed used by resulting in faster, more uniform and, therefore, reliable germination of these species. Finally, seeds of L. longifolia had low final germination (<40%), with a high presence of viable but dormant seeds. The ecological cues that promote germination in nature for both species should be further examined.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures
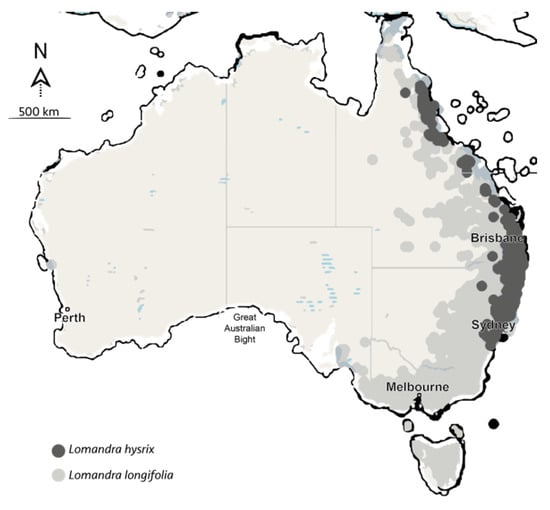
Figure 1
Open AccessArticle
Caffeic Acid and Biopesticides Interactions for the Control of Storage Beetles
by
, , , , , , , and
Appl. Biosci. 2023, 2(2), 211-221; https://doi.org/10.3390/applbiosci2020015 - 08 May 2023
Abstract
Infestations of stored-product pests cause significant losses of agricultural produce every year. Despite various environmental and health risks, chemical insecticides are now a ready-to-use solution for pest control. Against this background and in the context of Integrated Pest Management research, the present study
[...] Read more.
Infestations of stored-product pests cause significant losses of agricultural produce every year. Despite various environmental and health risks, chemical insecticides are now a ready-to-use solution for pest control. Against this background and in the context of Integrated Pest Management research, the present study focuses on the potential insecticidal effect of caffeic acid at five different concentrations (250, 500, 750, 1500 and 3000 ppm), and their combination with Cydia pomonella Granulovirus (CpGV), Bacillus thuringiensis subsp. tenebrionis and Beauveria bassiana strain GHA on three major insect stored-product beetle species, Tribolium confusum (Coleoptera: Tenebrionidae), Cryptolestes ferrugineus (Coleoptera: Laemophloeidae) and Trogoderma granarium Everts (Coleoptera: Dermestidae). Treatment efficacy was expressed as mortality in relation to exposure time and adult species number. Compared to the control, the results showed a clear dose-dependent pesticidal activity, expressed as significant adult mortality at a high-dose application, although some of the combinations of caffeic acid concentrations with the other substances acted positively (synergistically and additively) and some negatively. Based on our results, bioinsecticides can be combined with plant compounds such as caffeic acid and be integrated with other modern IPM tools in storage facilities.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
Open AccessFeature PaperReview
Crossiella, a Rare Actinomycetota Genus, Abundant in the Environment
by
, , , , , , , and
Appl. Biosci. 2023, 2(2), 194-210; https://doi.org/10.3390/applbiosci2020014 - 06 May 2023
Cited by 1
Abstract
The genus Crossiella contains two species, C. equi, causing nocardioform placentitis in horses, and C. cryophila, an environmental bacterium. Apart from C. equi, which is not discussed here, environmental Crossiella is rarely reported in the literature; thus, it has not
[...] Read more.
The genus Crossiella contains two species, C. equi, causing nocardioform placentitis in horses, and C. cryophila, an environmental bacterium. Apart from C. equi, which is not discussed here, environmental Crossiella is rarely reported in the literature; thus, it has not been included among “rare actinobacteria”, whose isolation frequency is very low. After C. cryophila, only five reports cover the isolation of Crossiella strains. However, the frequency of published papers on environmental Crossiella has increased significantly in recent years due to the extensive use of next-generation sequencing (NGS) and a huge cascade of data that has improved our understanding of how bacteria occur in the environment. In the last five years, Crossiella has been found in different environments (caves, soils, plant rhizospheres, building stones, etc.). The high abundance of Crossiella in cave moonmilk indicates that this genus may have an active role in moonmilk formation, as evidenced by the precipitation of calcite, witherite, and struvite in different culture media. This review provides an overview of environmental Crossiella, particularly in caves, and discusses its role in biomineralization processes and bioactive compound production.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biosciences 2023)
►▼
Show Figures
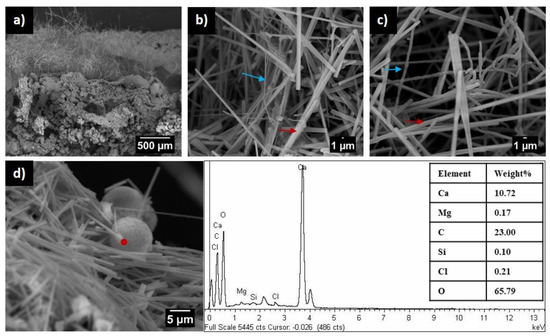
Figure 1
Open AccessArticle
Tissue Microarray Lipidomic Imaging Mass Spectrometry Method: Application to the Study of Alcohol-Related White Matter Neurodegeneration
Appl. Biosci. 2023, 2(2), 173-193; https://doi.org/10.3390/applbiosci2020013 - 04 Apr 2023
Abstract
Central nervous system (CNS) white matter pathologies accompany many diseases across the lifespan, yet their biochemical bases, mechanisms, and consequences have remained poorly understood due to the complexity of myelin lipid-based research. However, recent advances in matrix-assisted laser desorption/ionization-imaging mass spectrometry (MALDI-IMS) have
[...] Read more.
Central nervous system (CNS) white matter pathologies accompany many diseases across the lifespan, yet their biochemical bases, mechanisms, and consequences have remained poorly understood due to the complexity of myelin lipid-based research. However, recent advances in matrix-assisted laser desorption/ionization-imaging mass spectrometry (MALDI-IMS) have minimized or eliminated many technical challenges that previously limited progress in CNS disease-based lipidomic research. MALDI-IMS can be used for lipid identification, semi-quantification, and the refined interpretation of histopathology. The present work illustrates the use of tissue micro-arrays (TMAs) for MALDI-IMS analysis of frontal lobe white matter biochemical lipidomic pathology in an experimental rat model of chronic ethanol feeding. The use of TMAs combines workload efficiency with the robustness and uniformity of data acquisition. The methods described for generating TMAs enable simultaneous comparisons of lipid profiles across multiple samples under identical conditions. With the methods described, we demonstrate significant reductions in phosphatidylinositol and increases in phosphatidylcholine in the frontal white matter of chronic ethanol-fed rats. Together with the use of a novel rapid peak alignment protocol, this approach facilitates reliable inter- and intra-group comparisons of MALDI-IMS data from experimental models and could be extended to human disease states, including using archival specimens.
Full article
(This article belongs to the Special Issue Feature Papers for the Inaugural Issue of Applied Biosciences)
►▼
Show Figures
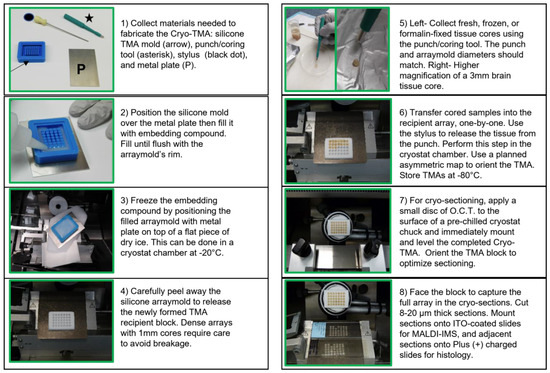

Figure 1
Open AccessReview
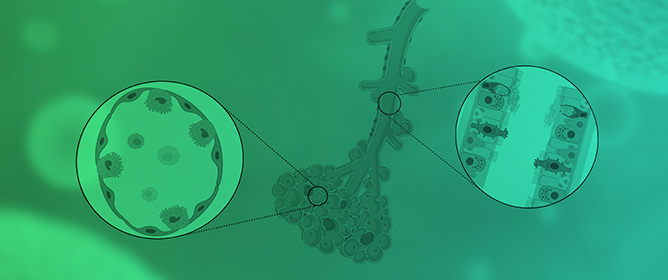
How Far Are We from Research That Is Independent of the Use of Animal Models? A Comparative Analysis between Animal and 3D/On-a-Chip Models for the Study of Respiratory Diseases
Appl. Biosci. 2023, 2(2), 157-172; https://doi.org/10.3390/applbiosci2020012 - 02 Apr 2023
Abstract
Over the last ten years, with the progress of in vitro culture methods, it has been possible to build increasingly reliable models to effectively mimic in vivo ones. The translational methodological approach that combined biotechnology and biomedical engineering has produced remarkable results, such
[...] Read more.
Over the last ten years, with the progress of in vitro culture methods, it has been possible to build increasingly reliable models to effectively mimic in vivo ones. The translational methodological approach that combined biotechnology and biomedical engineering has produced remarkable results, such as the development of ex vivo 3D culture models, the construction of on-a-chip organoids, and the construction of complex systems capable of bypassing the static nature of the two-dimensional cultural models that have been typical of in vitro studies conducted to date. However, nowadays, there is still reluctance to completely abandon the animal model as an essential reference or as an integrated step for the validation of a model or a proposed study. This is due to the partially correct conviction of the impossibility of reproducing, in vitro or ex vivo, the complexity of pathological models or the spatial communication between different cytotypes, as well as, more generally, the lack of systems capable of mimicking the dynamism of a complex in vivo system. In this study, we will compare different methodological approaches in the study of the three most common types of respiratory diseases: chronic obstructive pulmonary disease (COPD), asthma, and lung carcinomas. The purpose of this comparative study is to evaluate the most current methodological approaches to understand how far research is from being independent from animal models. Animal studies are generally considered necessary, but are still questioned because of the ethics and the cost–benefit ratio involved.
Full article
(This article belongs to the Special Issue Anatomy and Regenerative Medicine: From Methods to Applications)
►▼
Show Figures
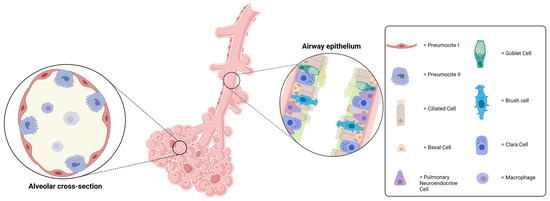
Figure 1
Open AccessReview
A Review of the Important Weapons against Antimicrobial Resistance in Sub-Saharan Africa
by
, , , and
Appl. Biosci. 2023, 2(2), 136-156; https://doi.org/10.3390/applbiosci2020011 - 01 Apr 2023
Cited by 1
Abstract
►▼
Show Figures
Antimicrobial resistance (AMR) is one of the top 10 global health threats facing humanity, and the sub-Saharan Africa (SSA) is among the heavily affected regions due to its weak health systems and limited resources. Due to an escalating number of AMR pathogens and
[...] Read more.
Antimicrobial resistance (AMR) is one of the top 10 global health threats facing humanity, and the sub-Saharan Africa (SSA) is among the heavily affected regions due to its weak health systems and limited resources. Due to an escalating number of AMR pathogens and the scarcity of new antimicrobials, efforts in the prevention of infections and the search for alternative treatment options are ongoing. The objective of this review was to assess important weapons against AMR in SSA. The highlighted weapons include vaccines, education and awareness, infection prevention and control (IPC) using water, sanitation, and hygiene (WASH), alternative treatment options, the One Health (OH) approach, AMR surveillance, operational national action plans (NAPs) on AMR, antimicrobial stewardship (AMS) programs, and good governance and regulations. Despite not being used at a satisfactory level in SSA, advanced techniques in dealing with AMR in SSA include (i) metagenomics, (ii) whole-genome sequencing (WGS) in AMR surveillance to track resistance trends and know when to intervene, and (iii) use of artificial intelligence in AMR prediction based on genomics data. The fight against AMR threat in SSA has embraced a number of currently available strategies, and developing new ones will lower the consequences of such a threat for future generations.
Full article
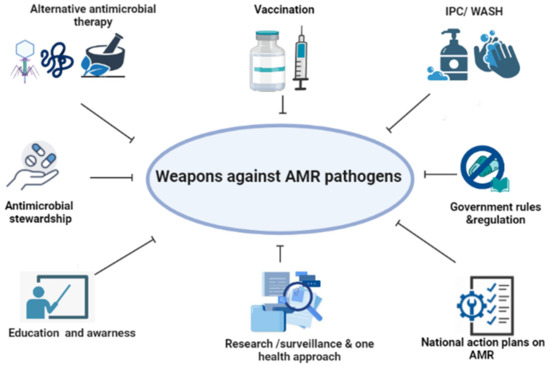
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Biosciences, Bioengineering, Biomolecules, JCDD, JCM, Micromachines, Reports
Usefulness and Clinical Applications of 3D Printing in Cardiovascular Diseases 2.0
Topic Editors: Zhonghua Sun, Massimo Chessa, Israel Valverde, Alexander Van De BruaeneDeadline: 31 October 2023
Topic in
Antioxidants, Applied Biosciences, Biomolecules, Horticulturae, Molecules, Pharmacy, Plants
Bioactive Substances, Pharmacognosy and Metabolomics
Topic Editors: Jiri Gruz, Lucie RárováDeadline: 10 December 2023
Topic in
Applied Biosciences, Bioengineering, Energies, Pollutants, Molecules
Sustainable Approaches for Biofuels from Waste Materials
Topic Editors: Md Sohrab Hossain, Venugopal BalakrishnanDeadline: 31 December 2023

Conferences
Special Issues
Special Issue in
Applied Biosciences
Plant Natural Compounds: From Discovery to Application
Guest Editors: Adriana Basile, Viviana Maresca, Piergiorgio CianciulloDeadline: 31 October 2023
Special Issue in
Applied Biosciences
Anatomy and Regenerative Medicine: From Methods to Applications
Guest Editors: Alessandro Pitruzzella, Alberto Fucarino, Fabio BucchieriDeadline: 30 November 2023
Special Issue in
Applied Biosciences
Feature Papers in Applied Biosciences 2023
Guest Editor: Robert HenryDeadline: 31 December 2023
Special Issue in
Applied Biosciences
The Role of Applied Bioscience in Pandemics
Guest Editors: Robert Henry, Burkhard Poeggeler, Nikolaos KourkoumelisDeadline: 31 January 2024















