Journal Description
Applied Microbiology
Applied Microbiology
is an international, peer-reviewed, open access journal on application of microorganisms published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the first half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
- Applied Microbiology is a companion journal of Microorganisms.
subject
Imprint Information
Open Access
ISSN: 2673-8007
Latest Articles
Safety and Effects of Intravaginal Administration of Lacticaseibacillus rhamnosus CRL1332 Immobilized on Nanofibers in a Murine Experimental Model
Appl. Microbiol. 2023, 3(3), 1013-1026; https://doi.org/10.3390/applmicrobiol3030069 - 24 Aug 2023
Abstract
The design of probiotic hygiene products for daily use is considered an adequate alternative for the restoration of the vaginal microbiome, maintaining health, and/or preventing infections of the female urogenital tract. Most of these probiotic products are available on the world market, but
[...] Read more.
The design of probiotic hygiene products for daily use is considered an adequate alternative for the restoration of the vaginal microbiome, maintaining health, and/or preventing infections of the female urogenital tract. Most of these probiotic products are available on the world market, but their efficacy and safety are not sufficiently documented. One of the requirements to transfer novel probiotic formulas/products to the productive sector is to demonstrate their innocuity and the absence of adverse or collateral effects on the host, mainly assayed in experimental models. The inclusion of beneficial lactobacilli in nanofibers by electrospinning technique has shown promising application possibilities, and the immobilization of Lacticaseibacillus rhamnosus CRL1332 in nanofibers with and without bioprotective substances and their characterization were previously performed by our research group. In this work, the safety of the intravaginal (i.va.) administration of these functional nanofibers in a murine experimental model was evaluated. L. rhamnosus CRL1332 immobilized in different nanofibers was intravaginally inoculated into mice (seven daily doses). Vaginal washes were taken for microbiological (cultivable lactobacilli) and cytological techniques, and the vagina was used for histological and morphological-ultrastructural evaluation. Our results demonstrated that the intravaginal administration of L. rhamnosus CRL1332 immobilized in nanofibers is safe in murine models, given the absence of an inflammatory response at the cytological and histological levels, with minor modifications at the ultrastructural level, and also related to the normal cultivable vaginal microbiota. On the other hand, the number of cultivable lactobacilli increased in the vagina of mice receiving L. rhamnosus CRL1332 nanofibers. The results indicate the safety of lactobacilli-functional nanofibers and support their inclusion in the design of vaginal probiotic products to prevent/treat urogenital infections and reconstitute the women’s vaginal microbiota.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►
Show Figures
Open AccessBrief Report
Selective Change in the Bacteria Cultured and Isolated in Respiratory Sputum from Elderly Patients during the SARS-CoV-2 Pandemic
Appl. Microbiol. 2023, 3(3), 1003-1012; https://doi.org/10.3390/applmicrobiol3030068 - 23 Aug 2023
Abstract
The SARS-CoV-2 pandemic has affected social patterns and consequently the prevalence of infections, such as seasonal influenza. It has been reported that invasive pneumococcal infection has markedly decreased worldwide. Method: We retrospectively investigated the bacteria cultured and isolated from 23,052 respiratory sputum samples
[...] Read more.
The SARS-CoV-2 pandemic has affected social patterns and consequently the prevalence of infections, such as seasonal influenza. It has been reported that invasive pneumococcal infection has markedly decreased worldwide. Method: We retrospectively investigated the bacteria cultured and isolated from 23,052 respiratory sputum samples obtained at our hospital from April 2015 to March 2022. The average patient age was 71.8 years old, with a standard deviation of 16.0 years old. There was no significant difference in the age of the patients or the female-to-male ratio between each year. The detection ratio of bacteria was analyzed in accordance with sputum quality based on the Geckler classification. Results: The detection ratio of community-acquired pneumonia pathogens such as Haemophilus influenzae, Moraxella catarrhalis, and Streptococcus pneumoniae increased in parallel with the quality of the sputum, while that of hospital-acquired pneumonia pathogens such as Klebsiella pneumoniae, Pseudomonas aeruginosa, and Staphylococcus aureus was not significantly affected by the quality of the sputum. The detection ratio of former pathogens in the good-quality respiratory sputum had decreased significantly since April 2020 by 60–80%, while that of P. aeruginosa and S. aureus had increased by 40–50%. Conclusions: The SARS-CoV-2 pandemic reduced the detection ratio of H. influenzae, M. catarrhalis, and S. pneumoniae but increased that of P. aeruginosa and S. aureus in the good-quality respiratory sputum from elderly patients. The influence of this selective change in isolated bacteria on the health and comorbidity of elderly patients remains to be investigated.
Full article
(This article belongs to the Special Issue Human Microbiota Influence on Human Health Status 2.0)
►▼
Show Figures
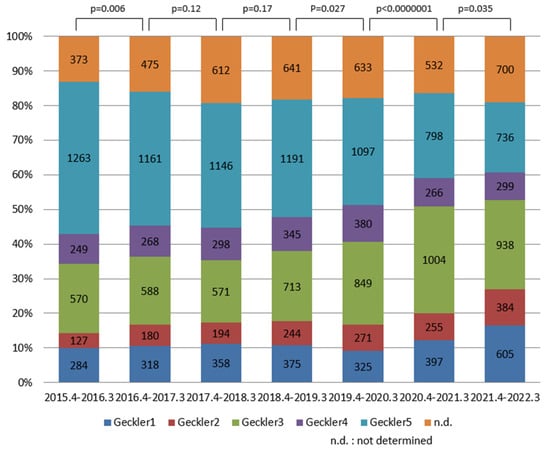
Figure 1
Open AccessSystematic Review
The Potential Role of Fecal Microbiota Transplantation in Parkinson’s Disease: A Systematic Literature Review
Appl. Microbiol. 2023, 3(3), 993-1002; https://doi.org/10.3390/applmicrobiol3030067 - 23 Aug 2023
Abstract
►▼
Show Figures
Background and Aims: Parkinson’s disease (PD) is a multifaceted disease that can cause symptoms in multiple body systems, including the gastrointestinal (GI) tract. Fecal microbiota transplantation (FMT) is a proposed treatment to address dysregulation in the microbiome, with neurologic and GI symptoms as
[...] Read more.
Background and Aims: Parkinson’s disease (PD) is a multifaceted disease that can cause symptoms in multiple body systems, including the gastrointestinal (GI) tract. Fecal microbiota transplantation (FMT) is a proposed treatment to address dysregulation in the microbiome, with neurologic and GI symptoms as theoretic effects. The complex relationship between gut dysbiosis and PD symptoms suggests that FMT may have a role in therapy. Methods: A total of 124 articles with information pertaining to PD and FMT were reviewed. PD adult patients with neuromotor or GI symptoms who received FMT with moderate levels of evidence were examined. Data using self-reporting symptom scales were compared at baseline and following FMT. Results: An overall improvement was seen in patients’ neuromotor or GI symptoms, when compared to the baseline after FMT. After FMT, the patients reported improved constipation and decreased time to stool. Overall satisfaction differed between groups receiving differing routes of FMT administration and the severity of their baseline symptoms. Conclusions: FMT shows promising utility for PD patients with neuromotor and/or GI symptoms. Our results are limited by need to evaluate routes of administration, the chronicity of inflammation, and acidity’s role in colonization and efficacy. FMT has been illustrated as a well-tolerated non-pharmacologic treatment method for PD with refractory symptoms.
Full article
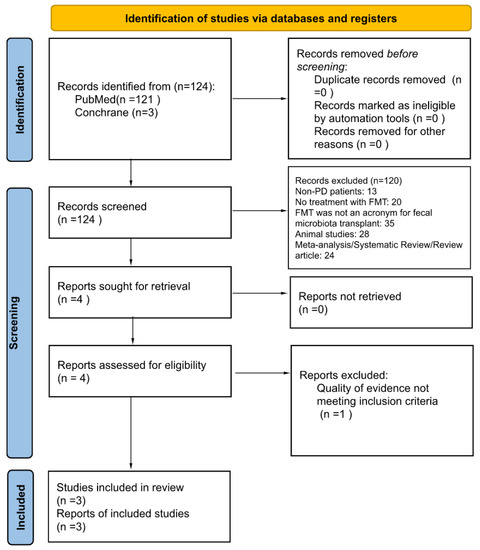
Figure 1
Open AccessArticle
Development of a Chicken Gastrointestinal Tract (GIT) Simulation Model: Impact of Cecal Inoculum Storage Preservation Conditions
by
, , , , , and
Appl. Microbiol. 2023, 3(3), 968-992; https://doi.org/10.3390/applmicrobiol3030066 - 22 Aug 2023
Abstract
A chicken gastrointestinal tract (GIT) simulation model was developed to help predict the potential effects of feed additives supplementation on chicken’ microbiota. The chemical and enzymatic conditions for oral, gastric, intestinal, and cecum fermentation phases were designed to closely resemble the chicken GIT
[...] Read more.
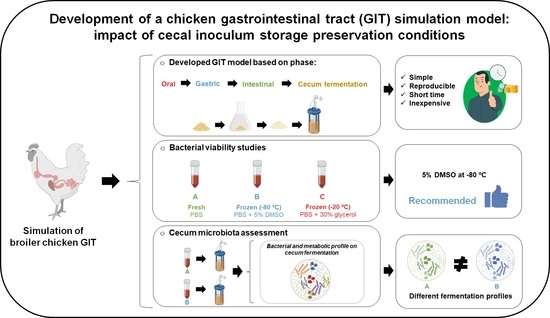
A chicken gastrointestinal tract (GIT) simulation model was developed to help predict the potential effects of feed additives supplementation on chicken’ microbiota. The chemical and enzymatic conditions for oral, gastric, intestinal, and cecum fermentation phases were designed to closely resemble the chicken GIT conditions. For cecum fermentation, the inoculum was obtained from the cecal contents of 18 38-day broiler chickens. The impact of inoculum preservation on bacteria viability was assessed by comparing two methods of preservation with fresh inoculum: (1) 5% dimethyl sulfoxide (DMSO) at −80 °C and (2) 30% glycerol at −20 °C. The fermentation with fresh and frozen (DMSO method) inoculums was performed and compared using standard chicken feed (SCF) and SCF with 1% fructooligosaccharides (FOS), and inoculum control (IC) condition without feed matrix was used as a baseline. Inoculum’s viability was assessed throughout 90 days of storage by culture media platting, while bacterial growth and metabolites production during fermentation was evaluated by quantitative polymerase chain reaction (qPCR), high-performance liquid chromatography (HPLC), and total ammonia nitrogen quantification. The DMSO method was shown to be the most suitable for cecal inoculum storage. Higher growth of beneficial cecal bacteria for fresh inoculum was observed in SCF while for frozen inoculum, was the SCF + FOS condition. Also, frozen inoculum had lower activity of butyrate producers and proteolytic bacteria, showing different fermentation profiles. The GIT model developed showed to be useful to test the effect of feed additives supplementation.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024))
►▼
Show Figures
Graphical abstract
Open AccessArticle
Screening of Rhizobacterial Isolates from Apple Rhizosphere for Their Biocontrol and Plant Growth Promotion Activity
by
, , , , , and
Appl. Microbiol. 2023, 3(3), 948-967; https://doi.org/10.3390/applmicrobiol3030065 - 22 Aug 2023
Abstract
Apple crops are prone to several diseases that limit their production—in particular, root rot caused by a new genus of oomycetes, mainly Phytopythium vexans. This study aims to screen antagonistic bacteria that can play an important role in the biological control of
[...] Read more.
Apple crops are prone to several diseases that limit their production—in particular, root rot caused by a new genus of oomycetes, mainly Phytopythium vexans. This study aims to screen antagonistic bacteria that can play an important role in the biological control of this pathogenic oomycete and to evaluate their capacity to promote plant growth. The dual culture test revealed that, out of 200 bacterial isolates, 16 have been able to inhibit the mycelial growth of P. vexans with inhibition rates greater than 50%. The selected isolates were identified based on the 16S rDNA genes: 14 bacteria belonging to the genus Bacillus, Stenotrophomonas, and the family Enterobacteriaceae. Notably, two isolates, B1 and M2-6 (identified as Bacillus velezensis), demonstrated the highest inhibition rates of 70% and 68%, respectively. These selected isolates were examined for their ability to produce different compounds related to biocontrol and plant growth promotion. Furthermore, the 16 selected isolates were evaluated for their ability to produce compounds associated with biocontrol and plant growth promotion, including hydrolytic enzymes (cellulases, proteases, and amylases), HCN (hydrogen cyanide) production, phosphate solubilization, IAA (indole-3-acetic acid) production, pectinase production, and stimulation of sorghum bicolor growth in vivo. Variations were observed among the bacterial isolates in terms of their compound production and phytostimulation capabilities. However, the secretion of proteases was consistently detected in all antagonistic isolates. The presence of genes responsible for the production of antifungal lipopeptides (bacillomycin, fengycin, and iturin) in the selected bacterial isolates was determined using polymerase chain reaction (PCR) techniques, while the absence of genes involved in surfactin biosynthesis was also confirmed through PCR studies. These isolates demonstrated inhibitory activity through the production of proteases and antifungal lipopeptides. Further research is needed to explore their potential use in biological control strategies and to improve apple crop productivity.
Full article
(This article belongs to the Special Issue Applied Microbiology of Foods 2.0)
►▼
Show Figures
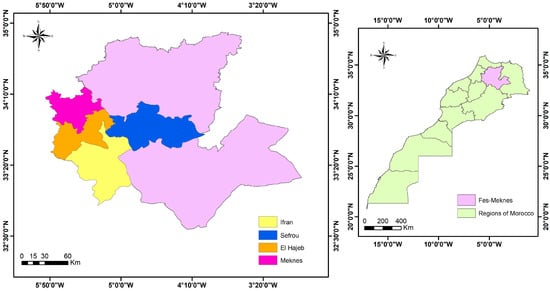
Figure 1
Open AccessReview
Overview of Probiotic Strains of Weizmannia coagulans, Previously Known as Bacillus coagulans, as Food Supplements and Their Use in Human Health
Appl. Microbiol. 2023, 3(3), 935-947; https://doi.org/10.3390/applmicrobiol3030064 - 21 Aug 2023
Abstract
Weizmannia coagulans, previously known as Bacillus coagulans and before that as Lactobacillus sporogenes, is a spore-forming, lactic acid-producing, Gram-positive, bacillus-shaped bacterial species with several known probiotic strains, including GBI-30, 6086 Unique IS-2, MTCC 5856, LBSC (DSM 17654), TBC169, SNZ 1969, BC30, and
[...] Read more.
Weizmannia coagulans, previously known as Bacillus coagulans and before that as Lactobacillus sporogenes, is a spore-forming, lactic acid-producing, Gram-positive, bacillus-shaped bacterial species with several known probiotic strains, including GBI-30, 6086 Unique IS-2, MTCC 5856, LBSC (DSM 17654), TBC169, SNZ 1969, BC30, and T11. This review focusses on the health benefit of these strains. A total of 53 clinical trials were found to use various strains of Weizmannia coagulans. However, 19 of these clinical trials did not provide strain information. Clinical evidence has shown that supplementation with strains of Weizmannia coagulans resulted in statistically significant health effects in the probiotic groups compared to the placebo. Several health benefits of the Weizmannia coagulans strains were found including relieving symptoms of irritable bowel syndrome, constipation, diarrhoea, and other gastrointestinal symptoms, function recovery treatment of non-fatty liver disease, after surgery or in patients with rheumatoid arthritis, quality of life and glucose- and lipid-related biomarkers related to overweight or obese participants or diabetic patients, absorption of protein or muscle integrity and improvement of peri- and post-menopausal symptoms. The main mechanism of action is the modulation of the intestinal microbiota and host immunity. However, in terms of several clinical studies involving small patient populations, others did not provide strain information. Larger, well-designed clinical studies are warranted to support the health benefits of Weizmannia coagulans strains.
Full article
Open AccessArticle
Response of Hypolimnetic Water and Bottom Sediment Microbial Communities to Freshwater Salinization—A Microcosm Experiment
Appl. Microbiol. 2023, 3(3), 915-934; https://doi.org/10.3390/applmicrobiol3030063 - 19 Aug 2023
Abstract
►▼
Show Figures
The introduction of NaCl in freshwater caused by winter runoffs is a problem whose consequences are still little understood. We sought to analyze the effect of NaCl addition on microbial communities of the hypolimnion and bottom sediments of a Canadian lake. Using microcosms
[...] Read more.
The introduction of NaCl in freshwater caused by winter runoffs is a problem whose consequences are still little understood. We sought to analyze the effect of NaCl addition on microbial communities of the hypolimnion and bottom sediments of a Canadian lake. Using microcosms comprising a salinity gradient varying between 0.01 and 3.22 ppt (10–3220 mg/L−1) NaCl, we investigated the effect of salinity on prokaryotic absolute abundance and diversity, following a three- and six-week exposure, and detected the presence of a salinity threshold for microbial communities’ differentiation. We observed a significant decline of bacterial diversity after six weeks in hypolimnetic samples. In the sediments, no clear effect of NaCl was observed on abundance or diversity, despite the presence of variations throughout the salinity gradient. The implication of nutrient fluctuations as well as the co-occurrence of species and inter-domain interactions is likely and would strongly contribute to the development of salt-exposed prokaryotic communities. In hypolimnetic water and sediments, the archaeal and eukaryotic communities differed significantly from 0.93 ppt (930 mg/L−1), while only conclusive at 1.9 ppt (1900 mg/L−1) NaCl in bacteria, meaning that the regulations in place are possibly suitable for the protection of the microbial communities in the hypolimnion and sediment lake layers.
Full article

Figure 1
Open AccessCommunication
Comparative Efficacy of Systemic and Combination Fungicides for the Control of Alternaria Leaf Spot of Cabbage
Appl. Microbiol. 2023, 3(3), 906-914; https://doi.org/10.3390/applmicrobiol3030062 - 12 Aug 2023
Abstract
Alternaria leaf spot of cabbage, caused by the Alternaria brassicicola, affects leaves of cabbages and often results in head rots causing severe decline in yield. In this work, the effects of systemic and combination fungicides on A. brassicicola mycelia growth in vitro
[...] Read more.
Alternaria leaf spot of cabbage, caused by the Alternaria brassicicola, affects leaves of cabbages and often results in head rots causing severe decline in yield. In this work, the effects of systemic and combination fungicides on A. brassicicola mycelia growth in vitro and disease severity in field trials were investigated. The results of in vitro evaluation revealed that both fungicides significantly inhibited (p < 0.05) the growth of A. brassicicola under in vitro conditions. However, metalaxyl-M 6% was less effective with 100 μg/mL having only 30 ± 3.5% inhibition. On the other hand, 100 μg/mL of mancozeb 63% + carbendazim 12% had 94 ± 3.5% growth inhibition of A. brassicicola, respectively, under the same conditions. Dose-response analysis of the efficacy of the two fungicides showed that the LC50 of metalaxyl-M 6% and mancozeb 63% + carbendazim 12% were 125.52 ppm and 57.22 ppm, respectively, indicating the superiority of combination fungicide over systemic fungicide alone. Field studies showed that while manure type significantly impacted on biomass production (p < 0.001), it did not significantly affect disease severity. On the other hand, the frequency of fungicide application impacted on disease severity, with biweekly application leading to a significant reduction in disease severity after 10 weeks.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
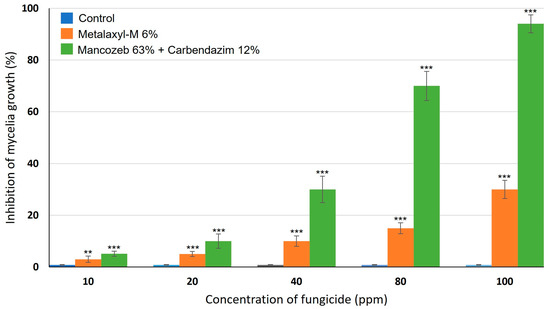
Figure 1
Open AccessReview
The Human Superorganism: Using Microbes for Freedom vs. Fear
Appl. Microbiol. 2023, 3(3), 883-905; https://doi.org/10.3390/applmicrobiol3030061 - 10 Aug 2023
Abstract
Balanced fear supports human rational decision-making and useful behavioral responses. In contrast, overwhelming, persistent, and unbalanced fear can paralyze the individual and result in heightened anxiety, lack of cognitive flexibility, fear-based public compliance and serious mental health issues. Psychobiotics research has established that
[...] Read more.
Balanced fear supports human rational decision-making and useful behavioral responses. In contrast, overwhelming, persistent, and unbalanced fear can paralyze the individual and result in heightened anxiety, lack of cognitive flexibility, fear-based public compliance and serious mental health issues. Psychobiotics research has established that a healthy microbiome is required for balanced fear and mental health protection via control of fear extinction. The recent COVID-19 pandemic featured daily, persistent, fear-of-a-single-contagion conditioning on a global scale paired with various behavioral mandates (e.g., lockdowns of the healthy, required wearing of face masks in many locations including schools, isolation from environmental microbes and each other through the closure of beaches and parks, and restrictions on social gatherings including access to family members in hospitals and senior-assisted facilities). Such mandates degraded the human microbiome and isolated us from each other and useful environmental microbes. It also ignored the historic role of secondary bacterial pathogens in pandemic deaths. This narrative review examines how the institutional promotion of fear-of-a-single-contagion, lack of balanced risk communication, and appalling disregard of our fundamental nature (as majority-microbial human superorganisms) resulted in problems rather than solutions. This review illustrates that government-public health-media promotion of pervasive fear and microbiome-degrading behaviors: (1) increased public compliance, (2) reduced cognitive flexibility, and (3) increased risk of mental health conditions. However, a portion of the general public chose a healthier path through their increased consumption of microbiome- and immune-supportive supplements and fermented foods during and after the COVID-19 pandemic. For a healthier future, public health must follow the lead of this population to ensure that human freedom, rather than paralyzing fear, dominates our future.
Full article
(This article belongs to the Special Issue Human Microbiota Influence on Human Health Status 2.0)
Open AccessArticle
Antibiotic Resistance Patterns of Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa Isolated from Hospital Wastewater
Appl. Microbiol. 2023, 3(3), 867-882; https://doi.org/10.3390/applmicrobiol3030060 - 08 Aug 2023
Abstract
►▼
Show Figures
Antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs) in treated hospital wastewater effluents constitute a major environmental and public health concern. The aim of this study was to investigate the antibiotic resistance patterns of Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa
[...] Read more.
Antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs) in treated hospital wastewater effluents constitute a major environmental and public health concern. The aim of this study was to investigate the antibiotic resistance patterns of Escherichia coli, Klebsiella pneumoniae, and Pseudomonas aeruginosa isolated from wastewater effluent at the Benjamin Mkapa Hospital (BMH) in Dodoma, Tanzania. These bacteria were selected to represent the most prevalent gram-negative bacteria found in hospital wastewater, and they have the potential to generate resistance and spread resistance genes to antibiotics. The wastewater BMH is treated in a Constructed Wetland (CW) planted with Typha latifolia before being released into the environment. The bacteria were isolated from wastewater effluent collected at the outlet of the CW. Isolated bacteria were analyzed for antibiotic resistance by disc diffusion method. Molecular identification of bacterial species was performed by using 16S rRNA. The results show that Klebsiella ssp. was the most common isolate detected, with a prevalence of 39.3%, followed by E. coli (27.9%) and Pseudomonas ssp. (18.0%). Klebsiella ssp. were more resistant than Pseudomonas ssp. for Tetracycline, Gentamycin, Ciprofloxacin, and Sulfamethoxazole. Pseudomonas ssp. were more resistant than Klebsiella ssp. for Ceftriaxone and Azithromycin. Klebsiella ssp. harbored more resistance genes (40%), followed by Pseudomonas ssp. (35%) and E. coli (20%). The findings of this investigation indicate that the effluent from the CW requires additional treatment to reduce discharged ARB and ARGs in the receiving water bodies. As a result, the effluent quality of the CW should be continuously monitored and assessed, and further developments for treating the final effluent are necessary.
Full article
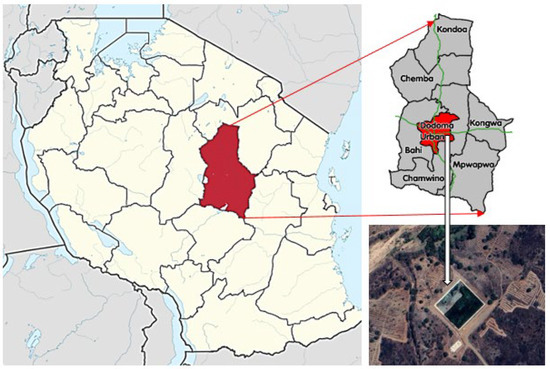
Figure 1
Open AccessReview
Reviewing the Current Understanding of Replant Syndrome in Orchards from a Soil Microbiome Perspective
Appl. Microbiol. 2023, 3(3), 856-866; https://doi.org/10.3390/applmicrobiol3030059 - 04 Aug 2023
Abstract
Replant syndrome (RS) of fruit and nut trees causes reduced tree vigor and crop productivity in orchard systems due to repeated plantings of closely related tree species. Although RS etiology has not been clearly defined, the causal agents are thought to be a
[...] Read more.
Replant syndrome (RS) of fruit and nut trees causes reduced tree vigor and crop productivity in orchard systems due to repeated plantings of closely related tree species. Although RS etiology has not been clearly defined, the causal agents are thought to be a complex of soil microorganisms combined with abiotic factors and susceptible tree genetics. Different soil disinfection techniques alleviate RS symptoms by reducing the loads of the deleterious microbiome; however, the positive effect on crop growth is temporary. The goals of this paper are: (1) to conceptualize the establishment of the syndrome from a microbiome perspective and (2) to propose sustainable solutions to develop a beneficial microbiome to inhibit the onset of RS.
Full article
(This article belongs to the Special Issue Microbiome in Ecosystem 2.0)
►▼
Show Figures

Figure 1
Open AccessArticle
The Basis for Variations in the Biofilm Formation by Different Salmonella Species and Subspecies: An In Vitro and In Silico Scoping Study
Appl. Microbiol. 2023, 3(3), 841-855; https://doi.org/10.3390/applmicrobiol3030058 - 03 Aug 2023
Abstract
This study examined whether the presence/absence of biofilm-associated genes may indicate the potential for differences in the biofilm formation among the Salmonella species/subspecies. We conducted an in vitro study on the biofilm formation by eighteen Salmonella strains of different species/subspecies. Strains belonging to
[...] Read more.
This study examined whether the presence/absence of biofilm-associated genes may indicate the potential for differences in the biofilm formation among the Salmonella species/subspecies. We conducted an in vitro study on the biofilm formation by eighteen Salmonella strains of different species/subspecies. Strains belonging to subspecies enterica were generally poorer biofilm formers than strains belonging to species bongori and subspecies arizonae, diarizonae, and indica. A broader in silico study was subsequently conducted. The presence/absence of 57 biofilm-associated genes was further investigated among 323 Salmonella whole genomes of various species/subspecies. The lpfE gene was present in in 88.2% of subspecies enterica but was absent in ~90.2–100% of other subspecies. The sirA gene was present in 11.8% of subspecies enterica and 2.9% of S. diarizonae genomes while absent in other species/subspecies. The lpfe gene and sirA gene in subspecies enterica negatively correlated with environmental biofilm formation. The csrB gene was present in 71.4% of the S. arizonae and 94.3% of S. diarizonae genomes but absent in other species/subspecies. The absence of csrB in subspecies enterica positively correlated with weaker environmental biofilm formation. This may contribute to subspecies arizonae and diarizonae being better biofilm formers.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
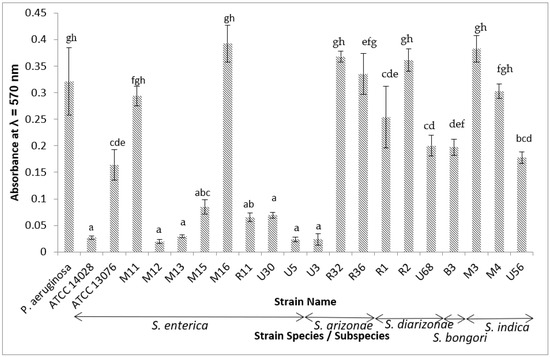
Figure 1
Open AccessArticle
Conversion of Unsaturated Short- to Medium-Chain Fatty Acids by Unspecific Peroxygenases (UPOs)
by
, , , , and
Appl. Microbiol. 2023, 3(3), 826-840; https://doi.org/10.3390/applmicrobiol3030057 - 19 Jul 2023
Abstract
Eighteen short- to medium-chain monounsaturated fatty acids were screened for hydroxylation and epoxidation using eleven different peroxygenase preparations. Most of these unspecific peroxygenases (UPOs) are secreted by fungal species of the dark-spored basidiomycetous families Psathyrellaceae and Strophariaceae, two belonged to the white-spored
[...] Read more.
Eighteen short- to medium-chain monounsaturated fatty acids were screened for hydroxylation and epoxidation using eleven different peroxygenase preparations. Most of these unspecific peroxygenases (UPOs) are secreted by fungal species of the dark-spored basidiomycetous families Psathyrellaceae and Strophariaceae, two belonged to the white-spored genus Marasmius (Marasmiaceae), and one belonged to the ascomycetous family Chaetomiaceae. The fatty acids (FAs) studied were categorized into three groups based on the position of the double bond: (i) terminal unsaturated FAs (between ω and ω-1), (ii) α-β-unsaturated FAs (between C2 and C3), and (iii) β-γ-unsaturated FAs (between C3 and C4). Their chain lengths ranged from three to nine carbon atoms. FAs with a terminal double bond were significantly oxidized by only two UPOs, namely CglUPO and CraUPO (peroxygenases from Chaetomium globosum and Coprinellus radians, respectively), producing different products. FAs with internal double bonds were converted by all tested UPOs. While epoxides were observed as products in the case of α-β-unsaturated fatty acids, only CglUPO formed β-γ-epoxides from the corresponding FAs. The product pattern of the other UPOs for β-γ-unsaturated FAs was quite similar. On the other hand, the product pattern for oxidized α-β-unsaturated FAs was more variable and, in some cases, specific to a particular UPO. For example, in the reaction with trans-2-nonenoic acid, the enzymes clustered into six groups based on the formed products.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
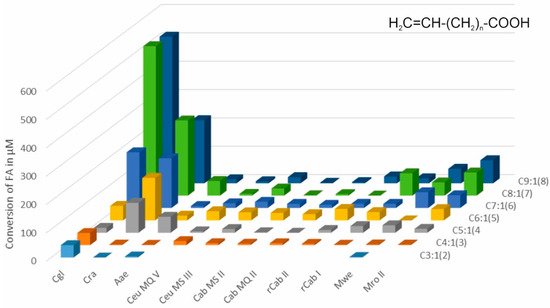
Figure 1
Open AccessReview
Yeast and Virus-like Particles: A Perfect or Imperfect Couple?
Appl. Microbiol. 2023, 3(3), 805-825; https://doi.org/10.3390/applmicrobiol3030056 - 14 Jul 2023
Abstract
Virus-like particles (VLPs) comprise viral structural proteins that self-assemble to form a particle similar to the native virus capsid. Since their discovery, they have been employed mainly as vaccines to prevent viral infection because they can elicit an immune response. Besides their use
[...] Read more.
Virus-like particles (VLPs) comprise viral structural proteins that self-assemble to form a particle similar to the native virus capsid. Since their discovery, they have been employed mainly as vaccines to prevent viral infection because they can elicit an immune response. Besides their use as vaccines, their application in cancer prevention and drug delivery is under intensive investigation. They can be produced in different systems such as bacteria, mammalian, plant, insect, and yeast cells. The main hurdle for their use is establishing a platform for production because many variables need to be considered. First, VLPs must be effective in the action for which they are constructed, depending on the nature of the VLPs. Second, the production platform must be suitable for safe and high-scale production. Yeast has been shown to be a valuable tool in VLP production, as it is able to express heterologous proteins efficiently and its manipulation is cheap and easy. Several species have been employed for this purpose. In the present review, we analyze the features of different yeast species and how they have been used to produce VLPs.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
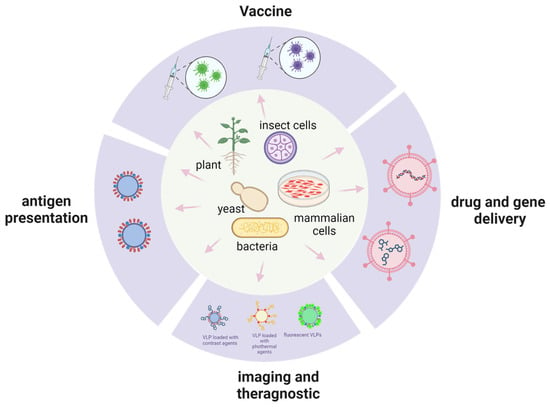
Figure 1
Open AccessReview
Perspectives on Using a Competitive Exclusion Approach to Control Listeria monocytogenes in Biological Soil Amendments of Animal Origin (BSAAO): A Review
Appl. Microbiol. 2023, 3(3), 786-804; https://doi.org/10.3390/applmicrobiol3030055 - 14 Jul 2023
Abstract
Biological soil amendments of animal origin (BSAAO), such as animal waste or animal-waste-based composts, may contain foodborne pathogens such as Listeria monocytogenes. Due to the ubiquitous nature of Listeria, it is essential to understand the behavior of L. monocytogenes in BSAAO
[...] Read more.
Biological soil amendments of animal origin (BSAAO), such as animal waste or animal-waste-based composts, may contain foodborne pathogens such as Listeria monocytogenes. Due to the ubiquitous nature of Listeria, it is essential to understand the behavior of L. monocytogenes in BSAAO in order to develop preharvest prevention strategies to reduce pathogen contamination. As biological control agents, competitive exclusion (CE) microorganisms have been widely utilized in agriculture to control plant- or foodborne pathogens. Due to the diverse microbial community, animal wastes and composts are the potential sources for isolating CE strains for pathogen control. To explore the potential of using CE to control L. monocytogenes in BSAAO, we thoroughly reviewed the studies on the fate of L. monocytogenes in the agriculture field, and in the isolation and identification of CE from different matrices, and the applications of CE as a biological control method. Future studies using a next-generation sequencing approach to identify and characterize CE strains in complex microbial communities can provide a comprehensive picture of the microbial interactions between invading pathogens and the indigenous microbiota in BSAAO. This comprehensive review will provide insight into the development of effective biological control measures for preventing L. monocytogenes contamination in the agricultural field and enhancing food safety.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
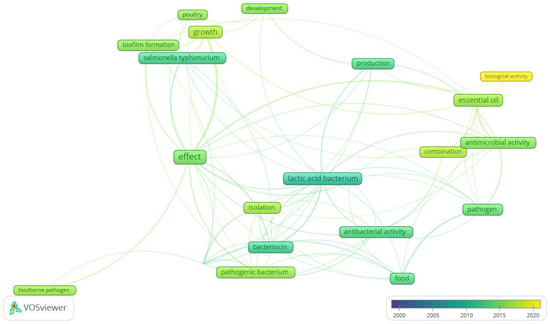
Figure 1
Open AccessArticle
The Probiotic Streptococcus salivarius M18 Increases Plasma Nitrite but Does Not Alter Blood Pressure: A Pilot Randomised Controlled Trial
by
, , , , and
Appl. Microbiol. 2023, 3(3), 774-785; https://doi.org/10.3390/applmicrobiol3030054 - 13 Jul 2023
Abstract
Some species of oral bacteria can reduce dietary nitrate to nitrite, which can later be converted to nitric oxide in the nitrate—nitrite—nitic oxide pathway. Increasing nitric oxide availability can reduce blood pressure (BP) and improve exercise performance. Streptococcus salivarius M18 (Streptococcus salivarius
[...] Read more.
Some species of oral bacteria can reduce dietary nitrate to nitrite, which can later be converted to nitric oxide in the nitrate—nitrite—nitic oxide pathway. Increasing nitric oxide availability can reduce blood pressure (BP) and improve exercise performance. Streptococcus salivarius M18 (Streptococcus salivarius M18) is a bacteriocin-producing probiotic that is known to improve oral health by inhibiting pathogenic oral bacteria. However, it is presently unclear whether probiotic-induced alterations to the oral microbiome will influence circulating levels of nitric oxide metabolites and BP. Purpose: To determine the effects of Streptococcus salivarius M18 supplementation on plasma and salivary nitrate and nitrite levels and BP. Methods: Ten healthy males (age 32 ± 8 y, body mass 88.2 ± 15.1 kg) completed 2 × 14-day supplementation phases in a randomized order at least 14 days apart. In one phase, participants consumed Streptococcus salivarius M18 probiotic lozenges (2.5 billion colony-forming units/dose) once per day, and in the other, they ingested water (placebo). The abundance of bacteria on the tongue was assessed via Illumina 16S rRNA gene sequencing, unstimulated saliva, and venous blood samples were collected, and BP was measured pre and post each phase. Saliva and plasma were analysed for nitrate and nitrite using chemiluminescence, and pH was measured in saliva. The change in each outcome from pre- to post-supplementation was compared between phases using repeated measures ANOVA. Results: Plasma nitrite increased from baseline following probiotic supplementation (from 173 ± 39 to 223 ± 63 nM, p = 0.003, 95% CI 192–250 nM). In comparison, there was no change in the placebo phase or between baselines (all p > 0.05). The abundance of nitrite-producing bacteria was not altered, salivary nitric oxide metabolites and pH did not change, and the increase in plasma nitrite did not result in reductions in BP (all p > 0.05). Conclusions: Supplementation with Streptococcus salivarius M18 increased plasma nitrite, a key marker of NO availability. Despite this, Streptococcus salivarius M18 did not lower BP in these healthy normotensive participants. Additionally, the increase in plasma nitrite was not associated with abundance changes in bacteria thought important to NO generation. Further research is required to determine the mechanism behind the increase in plasma nitrite and the potential therapeutic and ergogenic benefits of Streptococcus salivarius M18 supplementation.
Full article
(This article belongs to the Special Issue Human Microbiota Influence on Human Health Status 2.0)
►▼
Show Figures

Figure 1
Open AccessCommunication
A Snapshot of the Influent and Effluent Bacterial Populations in a Wastewater Treatment Plant in the North-West Province, South Africa
Appl. Microbiol. 2023, 3(3), 764-773; https://doi.org/10.3390/applmicrobiol3030053 - 13 Jul 2023
Abstract
►▼
Show Figures
Wastewater treatment plants receive influent wastewater that is contaminated with bacterial pathogens which may be released into the environment if the plant effluent is inadequately treated. In this study, next-generation sequencing was used to perform a 16S rDNA-based survey of bacterial populations in
[...] Read more.
Wastewater treatment plants receive influent wastewater that is contaminated with bacterial pathogens which may be released into the environment if the plant effluent is inadequately treated. In this study, next-generation sequencing was used to perform a 16S rDNA-based survey of bacterial populations in the influent and effluent from a treatment facility in the North-West Province (SA). In total, 3638 and 3872 effective DNA reads were obtained for the influent and effluent, respectively. Sequence analysis revealed the detection of a diverse bacterial constituency in both the influent and effluent samples. The phyla: Proteobacteria (49.82% and 52.04%), Firmicutes (14.06% and 13.14%) and Actinobacteria (5.00% and 9.99%) were found to be taxonomically abundant in the influent and effluent, respectively. This translated to the detection of biological treatment-, fecal coliform-, and disease-associated bacterial groups that are classified under the following genera: Escherichia spp., Serratia spp., Aeromonas spp., Legionella spp., Pseudomonas spp., Mycobacterium spp., Clostridium spp., Staphylococcus spp. and Streptococcus spp., Comamonas spp., Nitrosomonas spp., Acinetobacter spp., Rhodobacter spp., Paracoccus spp., Hyphomicrobium spp., and Desulfovibrio spp.
Full article
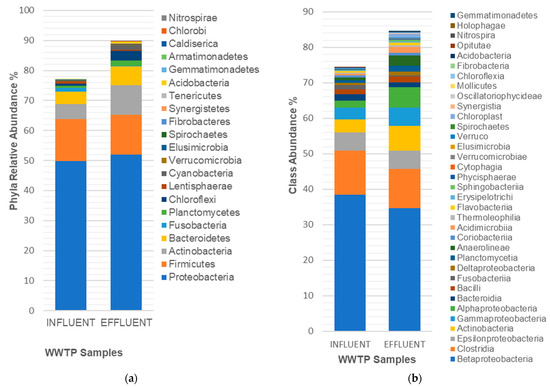
Figure 1
Open AccessArticle
Exploring the Feasibility of Intrapartum GBS Collection to Identify Residual GBS in a Pilot Study of an Antenatal Probiotic Intervention
Appl. Microbiol. 2023, 3(3), 752-763; https://doi.org/10.3390/applmicrobiol3030052 - 12 Jul 2023
Abstract
►▼
Show Figures
(1) Background: We aimed to explore the feasibility of collecting intrapartum maternal Group B Streptococcus (GBS) colonization and immediate post-birth neonatal GBS colonization cultures for use in a larger trial and to identify cases of residual GBS, which were hypothesized to be less
[...] Read more.
(1) Background: We aimed to explore the feasibility of collecting intrapartum maternal Group B Streptococcus (GBS) colonization and immediate post-birth neonatal GBS colonization cultures for use in a larger trial and to identify cases of residual GBS, which were hypothesized to be less common in the probiotics group. (2) Methods: This sub-study added additional outcome measures to the parent study to identify intrapartum and neonatal colonization and compare between probiotic and placebo groups and to identify cases of residual GBS. Intrapartum maternal vaginal and rectal GBS cultures were collected at the time of admission to a hospital for labor and to give birth. Neonatal oral and nasopharynx GBS cultures were collected within 1–2 h of giving birth. (3) Results: Thirty intrapartum samples were collected; twenty-eight had complete data. The antepartum GBS results significantly predicted the intrapartum results (p = 0.005), with 86.7% of cultures remaining the same at both time points. There were four cases where the intrapartum GBS results were different to the 36-week antepartum cultures results. A case of residual GBS was identified in one probiotic group participant. None of the neonatal swabs were positive for GBS. No cases of EOGBSD occurred in infants born to the study participants. (4) Conclusions: Although the 36–37 week GBS results significantly predicted the intrapartum results, the utility for a larger research trial on probiotics to reduce antenatal GBS is unclear. Intrapartum GBS swab collection was feasible in a busy nurse, midwife, and physician practice. GBS was not recovered from neonatal oral and nasopharyngeal swabs. The pathways of neonatal GBS colonization require further study.
Full article
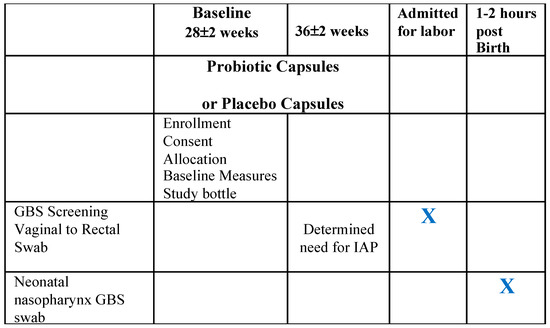
Figure 1
Open AccessReview
Fungal Pigments: Their Diversity, Chemistry, Food and Non-Food Applications
by
and
Appl. Microbiol. 2023, 3(3), 735-751; https://doi.org/10.3390/applmicrobiol3030051 - 12 Jul 2023
Abstract
Colorants have many applications in food, cosmetics, pharmaceutics, textile, paints, plastics, paper, ink and photographic industries. Colorants are classified according to their solubility into dyes and pigments. Those of natural origin have many advantages over synthetic ones, as natural colorants usually do not
[...] Read more.
Colorants have many applications in food, cosmetics, pharmaceutics, textile, paints, plastics, paper, ink and photographic industries. Colorants are classified according to their solubility into dyes and pigments. Those of natural origin have many advantages over synthetic ones, as natural colorants usually do not induce allergies or other health problems. In addition, their consumption in the food and drug industries is fortified with nutritional and health benefits as the majority of them possess antioxidant activity or can be used to produce some vitamins. Plants, animals, insects and microorganisms are rich sources of colorants. However, microbial pigments are favored over other natural pigments due to their higher yield, stability, economical production. Therefore, we focus in this review on fungal pigments, the history of their use, their chemistry and their applications in food and non-food fields. Additionally, the ability of the fungal genus, Epicoccum, to produce pigments is discussed. Moreover, the challenges and future prospects concerning fungal pigment production are highlighted in detail.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures

Figure 1
Open AccessArticle
Sugar-Induced Cell Death in the Yeast S. cerevisiae Is Accompanied by the Release of Octanoic Acid, Which Does Not Originate from the Fatty Acid Synthesis Type II Mitochondrial System
Appl. Microbiol. 2023, 3(3), 722-734; https://doi.org/10.3390/applmicrobiol3030050 - 12 Jul 2023
Abstract
Incubation of the yeast S. cerevisiae with glucose, in the absence of other nutrients, leads to Sugar-Induced Cell Death (SICD), accompanied by the accumulation of Reactive Oxygen Species (ROS). Yeast acidifies the environment during glucose metabolism not only as a result of the
[...] Read more.
Incubation of the yeast S. cerevisiae with glucose, in the absence of other nutrients, leads to Sugar-Induced Cell Death (SICD), accompanied by the accumulation of Reactive Oxygen Species (ROS). Yeast acidifies the environment during glucose metabolism not only as a result of the activity of the H+-ATPase of the plasma membrane but also due to the release of carboxylic acids. Acetic acid is known to induce apoptosis in growing yeast. We analyzed the composition of the incubation medium and found octanoic acid (OA) but no other carboxylic acids. Its concentration (0.675 µM) was significantly lower than the one at which OA had a toxic effect on the cell. However, the theoretically calculated concentration of OA inside the cell (about 200 μM) was found to be high enough to lead to cell necrosis. To test the hypothesis that OA might cause SICD, we used a ΔACP1 strain incapable of synthesizing OA in the yeast mitochondrial Fatty Acid Synthesis type II system (FAS-II). The deletion of the ACP1 gene did not affect the OA content in the medium. But, on the other hand, OA is a precursor of lipoic acid, which has antioxidant properties. However, strains with deleted genes for lipoic acid biosynthesis from OA (ΔPPT2, ΔLIP2, ΔLIP5, and ΔSGV3) showed no change in ROS and SICD levels. Thus, lipoic acid synthesized in FAS-II does not protect cells from ROS accumulated during SICD. We conclude that OA synthesized in the mitochondrial FAS-II system and its derivative lipoic acid are not involved in SICD in yeast S. cerevisiae.
Full article
(This article belongs to the Special Issue Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology)
►▼
Show Figures
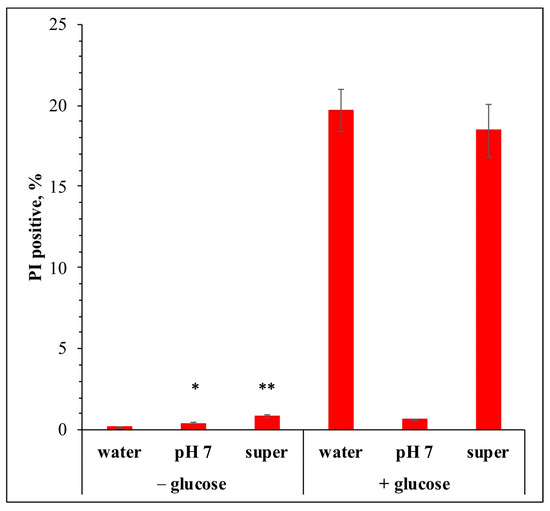
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Applied Microbiology, Biology, Biomedicines, Biomolecules, Proteomes
Biomedical Applications of Enzymes
Topic Editors: Amirata Saei Dibavar, Roozbeh Eskandari, Lucia CoppoDeadline: 30 October 2023
Topic in
Antibiotics, Applied Microbiology, Chemistry, Molecules, TropicalMed
Novel Antimicrobial Agents: Discovery, Design and New Therapeutic Strategies, 2nd Volume
Topic Editors: Mark G. Moloney, Sónia SilvaDeadline: 31 December 2023
Topic in
Applied Microbiology, Bioengineering, Biology, Environments, Microorganisms
Environmental Bioengineering and Geomicrobiology
Topic Editors: Xian-Chun Zeng, Deng LiuDeadline: 20 December 2024

Conferences
Special Issues
Special Issue in
Applied Microbiology
Human Microbiota Influence on Human Health Status 2.0
Guest Editor: Serena SchippaDeadline: 30 September 2023
Special Issue in
Applied Microbiology
Microbiome in Ecosystem 2.0
Guest Editor: Bong-Soo KimDeadline: 31 December 2023
Special Issue in
Applied Microbiology
Exclusive Papers Collection of Editorial Board Members and Invited Scholars in Applied Microbiology (2023, 2024)
Guest Editor: Ian ConnertonDeadline: 30 June 2024
Special Issue in
Applied Microbiology
Current Trends in the Applications of Probiotics and Other Beneficial Microbes
Guest Editor: Sabina FijanDeadline: 31 December 2024















