Journal Description
BioMedInformatics
BioMedInformatics
is an international, peer-reviewed, open access journal on all areas of biomedical informatics, as well as computational biology and medicine, published quarterly online by MDPI.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 16.6 days after submission; acceptance to publication is undertaken in 5.6 days (median values for papers published in this journal in the first half of 2023).
- Recognition of Reviewers: APC discount vouchers, optional signed peer review, and reviewer names published annually in the journal.
Latest Articles
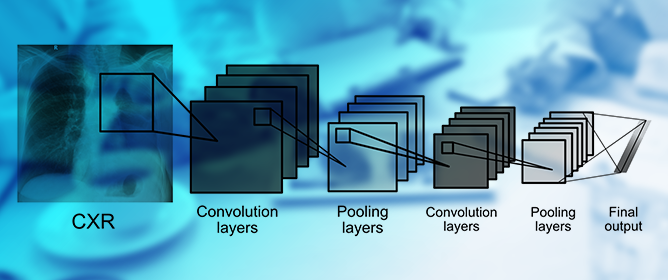
Automated Methods for Tuberculosis Detection/Diagnosis: A Literature Review
BioMedInformatics 2023, 3(3), 724-751; https://doi.org/10.3390/biomedinformatics3030047 (registering DOI) - 01 Sep 2023
Abstract
Tuberculosis (TB) is one of the leading infectious causes of death worldwide. The effective management and public health control of this disease depends on early detection and careful treatment monitoring. For many years, the microscopy-based analysis of sputum smears has been the most
[...] Read more.
Tuberculosis (TB) is one of the leading infectious causes of death worldwide. The effective management and public health control of this disease depends on early detection and careful treatment monitoring. For many years, the microscopy-based analysis of sputum smears has been the most common method to detect and quantify Mycobacterium tuberculosis (Mtb) bacteria. Nonetheless, this form of analysis is a challenging procedure since sputum examination can only be reliably performed by trained personnel with rigorous quality control systems in place. Additionally, it is affected by subjective judgement. Furthermore, although fluorescence-based sample staining methods have made the procedure easier in recent years, the microscopic examination of sputum is a time-consuming operation. Over the past two decades, attempts have been made to automate this practice. Most approaches have focused on establishing an automated method of diagnosis, while others have centred on measuring the bacterial load or detecting and localising Mtb cells for further research on the phenotypic characteristics of their morphology. The literature has incorporated machine learning (ML) and computer vision approaches as part of the methodology to achieve these goals. In this review, we first gathered publicly available TB sputum smear microscopy image sets and analysed the disparities in these datasets. Thereafter, we analysed the most common evaluation metrics used to assess the efficacy of each method in its particular field. Finally, we generated comprehensive summaries of prior work on ML and deep learning (DL) methods for automated TB detection, including a review of their limitations.
Full article
(This article belongs to the Section Clinical Informatics)
►
Show Figures
Open AccessFeature PaperArticle
Minimal Hip Joint Space Width Measured on X-rays by an Artificial Intelligence Algorithm—A Study of Reliability and Agreement
by
, , , , , , , and
BioMedInformatics 2023, 3(3), 714-723; https://doi.org/10.3390/biomedinformatics3030046 (registering DOI) - 01 Sep 2023
Abstract
Minimal joint space width (mJSW) is a radiographic measurement used in the diagnosis of hip osteoarthritis. A large variance when measuring mJSW highlights the need for a supporting diagnostic tool. This study aimed to estimate the reliability of a deep learning algorithm designed
[...] Read more.
Minimal joint space width (mJSW) is a radiographic measurement used in the diagnosis of hip osteoarthritis. A large variance when measuring mJSW highlights the need for a supporting diagnostic tool. This study aimed to estimate the reliability of a deep learning algorithm designed to measure the mJSW in pelvic radiographs and to estimate agreement between the algorithm and orthopedic surgeons, radiologists, and a reporting radiographer. The algorithm was highly consistent when measuring mJSW with a mean difference at 0.00. Human readers, however, were subject to variance with a repeatability coefficient of up to 1.31. Statistically, although not clinically significant, differences were found between the algorithm’s and all readers’ measurements with mean measured differences ranging from −0.78 to −0.36 mm. In conclusion, the algorithm was highly reliable, and the mean measured difference between the human readers combined and the algorithm was low, i.e., −0.5 mm bilaterally. Given the consistency of the algorithm, it may be a useful tool for monitoring hip osteoarthritis.
Full article
(This article belongs to the Section Imaging Informatics)
►▼
Show Figures
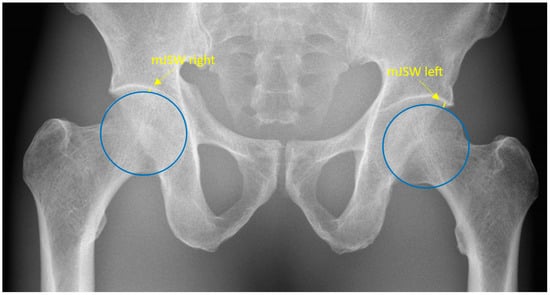
Figure 1
Open AccessReview
Deep Learning and Federated Learning for Screening COVID-19: A Review
BioMedInformatics 2023, 3(3), 691-713; https://doi.org/10.3390/biomedinformatics3030045 (registering DOI) - 01 Sep 2023
Abstract
Since December 2019, a novel coronavirus disease (COVID-19) has infected millions of individuals. This paper conducts a thorough study of the use of deep learning (DL) and federated learning (FL) approaches to COVID-19 screening. To begin, an evaluation of research articles published between
[...] Read more.
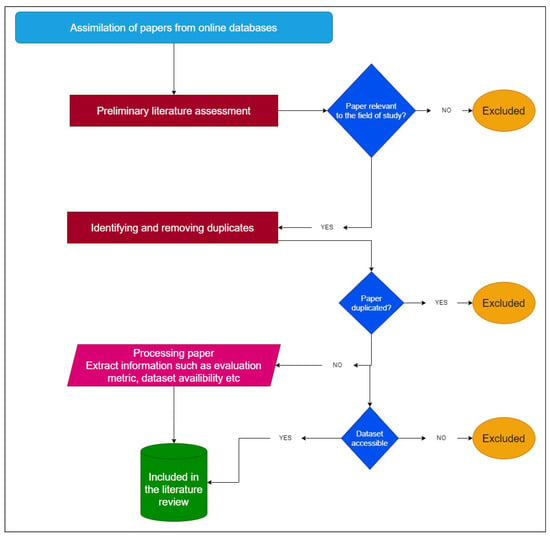
Since December 2019, a novel coronavirus disease (COVID-19) has infected millions of individuals. This paper conducts a thorough study of the use of deep learning (DL) and federated learning (FL) approaches to COVID-19 screening. To begin, an evaluation of research articles published between 1 January 2020 and 28 June 2023 is presented, considering the preferred reporting items of systematic reviews and meta-analysis (PRISMA) guidelines. The review compares various datasets on medical imaging, including X-ray, computed tomography (CT) scans, and ultrasound images, in terms of the number of images, COVID-19 samples, and classes in the datasets. Following that, a description of existing DL algorithms applied to various datasets is offered. Additionally, a summary of recent work on FL for COVID-19 screening is provided. Efforts to improve the quality of FL models are comprehensively reviewed and objectively evaluated.
Full article
(This article belongs to the Special Issue Features of Bioinformatic Analyses for SARS-CoV-2 Infections and Vaccination)
►▼
Show Figures
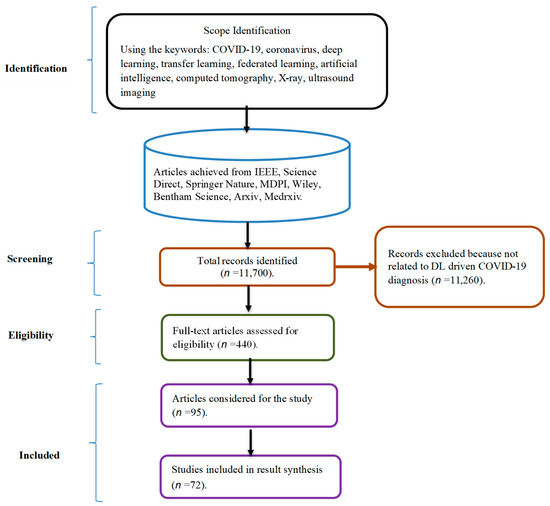
Figure 1
Open AccessArticle
Human Gm, Km, and Am Allotypes: WHO/IMGT Nomenclature and IMGT Unique Numbering for Immunoinformatics and Therapeutical Antibodies
BioMedInformatics 2023, 3(3), 649-690; https://doi.org/10.3390/biomedinformatics3030044 - 09 Aug 2023
Abstract
Human immunoglobulin allotypes are allelic antigenic determinants (or “markers”) determined serologically, classically by hemagglutination inhibition, on the human immunoglobulin (IG) or antibody heavy and light chains. The allotypes have been identified on the gamma1, gamma2, gamma3, and alpha2 heavy chains (designated as G1m,
[...] Read more.
Human immunoglobulin allotypes are allelic antigenic determinants (or “markers”) determined serologically, classically by hemagglutination inhibition, on the human immunoglobulin (IG) or antibody heavy and light chains. The allotypes have been identified on the gamma1, gamma2, gamma3, and alpha2 heavy chains (designated as G1m, G2m, G3m, and A2m allotypes, respectively) and on the kappa light chain (Km allotypes). Gm and Am allotypes have been one of the most powerful tools in population genetics, as they are inherited in fixed combinations, or Gm–Am haplotypes, owing to the linkage of the human IGHC genes in the IGH locus on chromosome 14. They have been very instrumental in molecular characterization of the human IGHC genes (gene polymorphisms or alleles, and IG heavy-chain structure in domains) and of the IGH locus (IGHC gene order, gene conversion, and copy number variation (CNV)). They represent a major system for understanding immunogenicity of the polymorphic IG chains in relation to amino acid and conformational changes. The WHO/IMGT allotype nomenclature and the IMGT unique numbering for constant (C) domain bridge Gm–Am and Km alleles to IGHC and IGKC gene alleles and structures and, by definition, to IG chain immunogenicity, opening the way for immunoinformatics of personalized therapeutic antibodies and engineered variants.
Full article
(This article belongs to the Section Applied Biomedical Data Science)
►▼
Show Figures
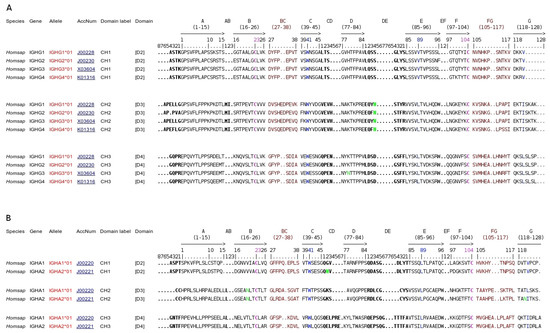

Figure 1
Open AccessArticle
Deployment of an Automated Method Verification-Graphical User Interface (MV-GUI) Software
BioMedInformatics 2023, 3(3), 632-648; https://doi.org/10.3390/biomedinformatics3030043 - 02 Aug 2023
Abstract
Clinical laboratories frequently conduct method verification studies to ensure that the process meets quality standards for its intended use, such as patient testing. They play a pivotal role in healthcare, but issues such as accurate statistical assessment and reporting of verification data often
[...] Read more.
Clinical laboratories frequently conduct method verification studies to ensure that the process meets quality standards for its intended use, such as patient testing. They play a pivotal role in healthcare, but issues such as accurate statistical assessment and reporting of verification data often make these studies challenging. Missteps can lead to false conclusions about method performance, risking patient safety or leading to incorrect diagnoses. Despite a requirement for accredited labs to document method performance, existing solutions are often expensive and complex. Addressing these issues, we present Method Verification-Graphical User Interface (MV-GUI), a software package designed for ease of use. It is platform-independent, capable of statistical analysis, and generates accreditation-ready reports swiftly and efficiently. Users can input patient data from one or more .CSV files, and MV-GUI will produce comprehensive reports, including statistical comparison tables, regression plots, and Bland–Altman plots. While method validation, which establishes the performance of new diagnostic tools, remains a crucial concern for manufacturers, MV-GUI primarily streamlines the method verification process. The software aids both medical practitioners and researchers and is designed to be user-friendly, even for non-experienced users. Requiring no internet connection, MV-GUI can operate in restricted IT environments, making method verification widely accessible and efficient.
Full article
(This article belongs to the Section Clinical Informatics)
►▼
Show Figures
Figure 1
Open AccessArticle
Effective Feature Engineering and Classification of Breast Cancer Diagnosis: A Comparative Study
BioMedInformatics 2023, 3(3), 616-631; https://doi.org/10.3390/biomedinformatics3030042 - 02 Aug 2023
Abstract
Breast cancer is among the most common cancers found in women, causing cancer-related deaths and making it a severe public health issue. Early prediction of breast cancer can increase the chances of survival and promote early medical treatment. Moreover, the accurate classification of
[...] Read more.
Breast cancer is among the most common cancers found in women, causing cancer-related deaths and making it a severe public health issue. Early prediction of breast cancer can increase the chances of survival and promote early medical treatment. Moreover, the accurate classification of benign cases can prevent cancer patients from undergoing unnecessary treatments. Therefore, the accurate and early diagnosis of breast cancer and the classification into benign or malignant classes are much-needed research topics. This paper presents an effective feature engineering method to extract and modify features from data and the effects on different classifiers using the Wisconsin Breast Cancer Diagnosis Dataset. We then use the feature to compare six popular machine-learning models for classification. The models compared were Logistic Regression, Random Forest, Decision Tree, K-Neighbors, Multi-Layer Perception (MLP), and XGBoost. The results showed that the Decision Tree model, when applied to the proposed feature engineering, was the best performing, achieving an average accuracy of 98.64%.
Full article
(This article belongs to the Special Issue Feature Papers in Computational Biology and Medicine)
►▼
Show Figures
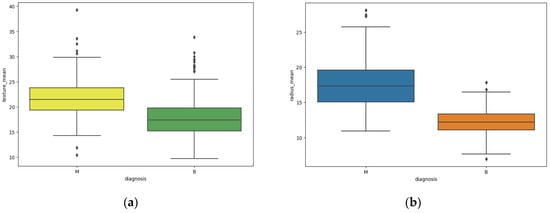
Figure 1
Open AccessArticle
Clinical and Demographic Attributes of Patients with Diabetes Associated with the Utilization of Telemedicine in an Urban Medically Underserved Population Area
BioMedInformatics 2023, 3(3), 605-615; https://doi.org/10.3390/biomedinformatics3030041 - 01 Aug 2023
Abstract
Marginalized populations often experience health disparities due to the significant obstacles to care associated with social, economic, and environmental inequities. When compared with advantaged social groups, these populations frequently experience increased risks, poorer health outcomes, and reduced quality of life (QoL). This research
[...] Read more.
Marginalized populations often experience health disparities due to the significant obstacles to care associated with social, economic, and environmental inequities. When compared with advantaged social groups, these populations frequently experience increased risks, poorer health outcomes, and reduced quality of life (QoL). This research examines the clinical and demographic characteristics—age, gender, and race—related to patients with varying stages of type 2 diabetes mellitus (T2DM), comparing the utilization of telemedicine (TM) with traditional healthcare face-to-face (F2F) appointments in an urban medically underserved population area (UMUPA). A logistic regression model, was used to analyze retrospective electronic patient health records (EHRs) from 1 January 2019 to 30 June 2021 of 265 patients with T2DM who had 3357 healthcare appointments. The overall percentage of healthcare provider appointments using TM was 46.7%, in comparison with 53.3% traditional F2F visits. Compared to patients with prediabetes, those with uncontrolled diabetes were more likely to utilize the TM mode of care rather than the traditional F2F mode (adjusted odds ratio (AoR), 1.33; confidence interval (CI), 1.07 to 1.64) after controlling for the other covariates in the model. Compared to patients in the age group 20–49 years, those in the age groups 50–64 years and ≥65 years had significantly lower odds (AoR, 0.78; CI, 0.65 to 0.94 and AoR, 0.71; CI, 0.58 to 0.88, respectively) of utilization of TM than the traditional F2F mode of care. White patients had significantly higher odds of using telemedicine rather than the traditional F2F mode (AoR, 1.25; CI, 1.07 to 1.47) when compared to the Black patients. Gender differences did not exist in the care utilization mode. As healthcare and public health continue to strive for health equity by eliminating health disparities within marginalized populations, it is essential that the mode of care for patients, such as those with T2DM, must evolve and adapt to the needs and resources of the patients. Multisectoral partners have the opportunity to employ a systems thinking approach to improve the technological elements related to the global health disparities crisis. An essential goal is to to create a user-friendly interface that prioritizes easy navigation, affordability, and accessiblity for populations in medically underserved regions to improve overall population health outcomes.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
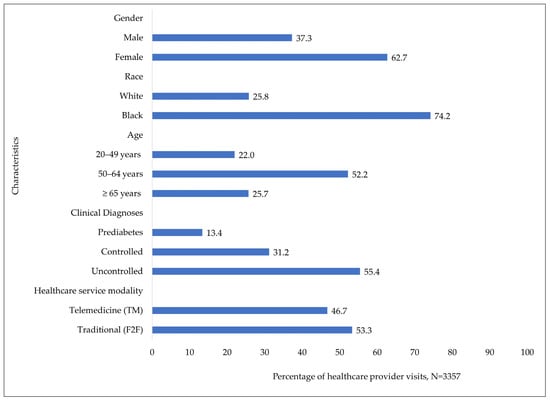
Figure 1
Open AccessArticle
A Machine Learning Pipeline for Cancer Detection on Microarray Data: The Role of Feature Discretization and Feature Selection
BioMedInformatics 2023, 3(3), 585-604; https://doi.org/10.3390/biomedinformatics3030040 - 01 Aug 2023
Abstract
Early disease detection using microarray data is vital for prompt and efficient treatment. However, the intricate nature of these data and the ongoing need for more precise interpretation techniques make it a persistently active research field. Numerous gene expression datasets are publicly available,
[...] Read more.
Early disease detection using microarray data is vital for prompt and efficient treatment. However, the intricate nature of these data and the ongoing need for more precise interpretation techniques make it a persistently active research field. Numerous gene expression datasets are publicly available, containing microarray data that reflect the activation status of thousands of genes in patients who may have a specific disease. These datasets encompass a vast number of genes, resulting in high-dimensional feature vectors that present significant challenges for human analysis. Consequently, pinpointing the genes frequently associated with a particular disease becomes a crucial task. In this paper, we present a method capable of determining the frequency with which a gene (feature) is selected for the classification of a specific disease, by incorporating feature discretization and selection techniques into a machine learning pipeline. The experimental results demonstrate high accuracy and a low false negative rate, while significantly reducing the data’s dimensionality in the process. The resulting subsets of genes are manageable for clinical experts, enabling them to verify the presence of a given disease.
Full article
(This article belongs to the Special Issue Feature Papers in Applied Biomedical Data Science)
►▼
Show Figures
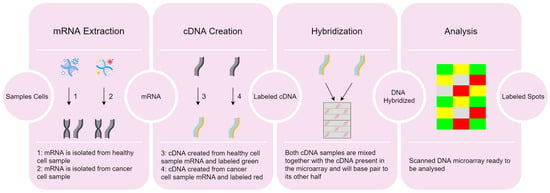
Figure 1
Open AccessReview
A Comprehensive Survey of Digital Twins in Healthcare in the Era of Metaverse
by
and
BioMedInformatics 2023, 3(3), 563-584; https://doi.org/10.3390/biomedinformatics3030039 - 21 Jul 2023
Cited by 1
Abstract
Digital twins (DTs) are becoming increasingly popular in various industries, and their potential for healthcare in the metaverse continues to attract attention. The metaverse is a virtual world where individuals interact with digital replicas of themselves and the environment. This paper focuses on
[...] Read more.
Digital twins (DTs) are becoming increasingly popular in various industries, and their potential for healthcare in the metaverse continues to attract attention. The metaverse is a virtual world where individuals interact with digital replicas of themselves and the environment. This paper focuses on personalized and precise medicine and examines the current application of DTs in healthcare within the metaverse. Healthcare practitioners may use immersive virtual worlds to replicate medical scenarios, improve teaching experiences, and provide personalized care to patients. However, the integration of DTs in the metaverse poses technical, regulatory, and ethical challenges that need to be addressed, including data privacy, standards, and accessibility. Through this examination, we aim to provide insights into the transformative potential of DTs in healthcare within the metaverse and encourage further research and development in this exciting domain.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Evaluation of Replies to Voice Queries in Gynecologic Oncology by Virtual Assistants Siri, Alexa, Google, and Cortana
by
, , , , , , , , , and
BioMedInformatics 2023, 3(3), 553-562; https://doi.org/10.3390/biomedinformatics3030038 - 11 Jul 2023
Abstract
Women that receive news that they have a malignancy of gynecologic origin can have questions about their diagnosis. These questions might be posed as voice queries to the virtual assistants Siri, Alexa, Google, and Cortana. Because our world has increasingly adopted smart phones
[...] Read more.
Women that receive news that they have a malignancy of gynecologic origin can have questions about their diagnosis. These questions might be posed as voice queries to the virtual assistants Siri, Alexa, Google, and Cortana. Because our world has increasingly adopted smart phones and standalone voice query devices, this study focused on the accuracy of audible replies by the virtual assistants (VAs) Siri, Alexa, Google, and Cortana to voice queries related to gynecologic oncology. Twenty-one evaluators analyzed VA audible answers to select voice queries related to gynecologic oncology. Questions were posed in three different ways for each voice query in order to maximize the likelihood of acceptability to the VAs in a 24-question panel. For general queries that were not related to gynecologic oncology, Google provided the most correct audible replies (83.3% correct), followed by Alexa (66.7% correct), Siri (45.8% correct), and Cortana (20.8% correct). For gynecologic oncology-related queries, the accuracy of the VAs was considerably lower: Google provided the most correct audible replies (18.1%), followed by Alexa (6.5%), Siri (5.5%), and Cortana (2.3%). There was a considerable drop in the accuracy of audible replies to oral queries on topics in gynecologic oncology relative to general queries that were not related to gynecologic oncology. There is considerable room for improvement in VA performance, so that caution is advised when using VAs for medical queries in gynecologic oncology. Our specific findings related to gynecologic oncology extend the work of others with regard to the low usability of general medical information obtained from VAs, so that reliance on conversational assistants for actionable medical information represents a safety risk for patients and consumers.
Full article
(This article belongs to the Special Issue Computational Biology and Artificial Intelligence in Medicine)
►▼
Show Figures

Figure 1
Open AccessData Descriptor
NJN: A Dataset for the Normal and Jaundiced Newborns
BioMedInformatics 2023, 3(3), 543-552; https://doi.org/10.3390/biomedinformatics3030037 - 05 Jul 2023
Abstract
Neonatal jaundice is a prevalent condition among newborns, with potentially severe complications that can result in permanent brain damage if left untreated during its early stages. The existing approaches for jaundice detection involve invasive procedures such as blood sample collection, which can inflict
[...] Read more.
Neonatal jaundice is a prevalent condition among newborns, with potentially severe complications that can result in permanent brain damage if left untreated during its early stages. The existing approaches for jaundice detection involve invasive procedures such as blood sample collection, which can inflict pain and distress on the patient, and may give rise to additional complications. Alternatively, a non-invasive method using image-processing techniques and implementing kNN, Random Forest, and XGBoost machine learning algorithms as a classifier can be employed to diagnose jaundice, necessitating a comprehensive database of infant images to achieve a diagnosis with high accuracy. This data article presents the NJN collection, a repository of newborn images encompassing diverse birthweights and skin tones, spanning an age range of 2 to 8 days. The dataset is accompanied by an Excel sheet file in CSV format containing the RGB and YCrCb channel values, as well as the status of each sample. The dataset and associated resources are openly accessible at Zenodo website. Moreover, the Python code for data testing utilizing various AI techniques is provided. Consequently, this article offers an unparalleled resource for AI researchers, enabling them to train their AI systems and develop algorithms that can assist neonatal intensive care unit (NICU) healthcare specialists in monitoring neonates while facilitating the fast, real-time, non-invasive, and accurate diagnosis of jaundice.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
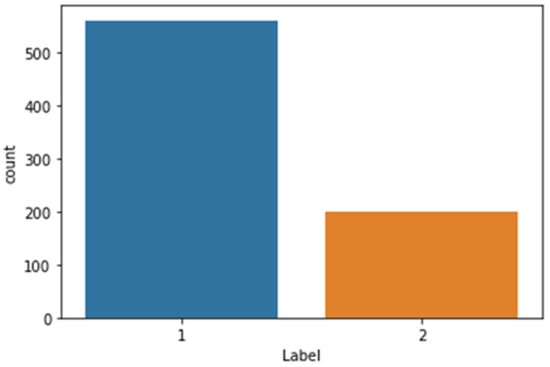
Figure 1
Open AccessArticle
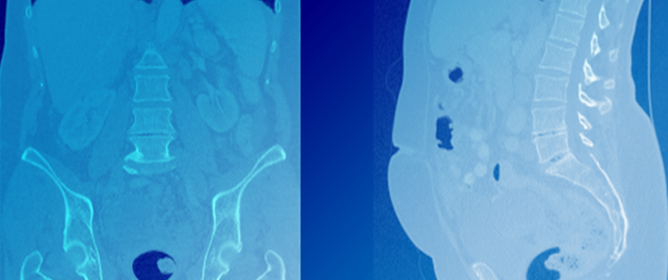
Improvement in Disease Diagnosis in Computed Tomography Images by Correlating Organ Volumes with Disease Occurrences in Humans
BioMedInformatics 2023, 3(3), 526-542; https://doi.org/10.3390/biomedinformatics3030036 - 05 Jul 2023
Abstract
Some diseases are known to cause or coincide with volume changes of certain structures in the body. Since these changes can be used to identify diseases, in this paper, we aimed to discover such new correlations. To this end, we trained a machine
[...] Read more.
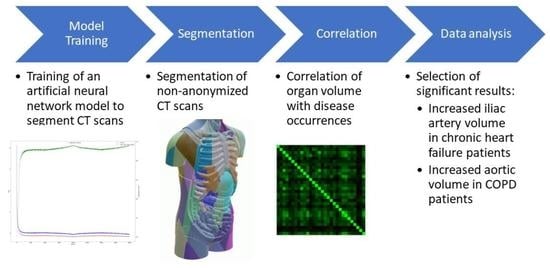
Some diseases are known to cause or coincide with volume changes of certain structures in the body. Since these changes can be used to identify diseases, in this paper, we aimed to discover such new correlations. To this end, we trained a machine learning model based on the TotalSegmentator model on computed tomography (CT) image data, to segment 104 anatomical structures, while trying to improve the accuracy of the model. We then used the model to segment CT scans of decedents who had at least one of 18 diseases. After correlating the structure volumes with disease occurrences, a possible new correlation between chronic artery failure and iliac artery volume was found and others were confirmed. However, due to the limitations of the model and the underlying data, further research is required.
Full article
(This article belongs to the Section Imaging Informatics)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Using Machine Learning to Expand the Ann Arbor Staging System for Hodgkin and Non-Hodgkin Lymphoma
BioMedInformatics 2023, 3(3), 514-525; https://doi.org/10.3390/biomedinformatics3030035 - 03 Jul 2023
Abstract
The Ann Arbor system is disadvantaged in utilizing information from additional prognostic factors. In this study, we applied the Ensemble Algorithm for Clustering Cancer Data (EACCD) to create a prognostic system for lymphoma that integrates additional prognostic factors. Hodgkin and non-Hodgkin lymphoma survival
[...] Read more.
The Ann Arbor system is disadvantaged in utilizing information from additional prognostic factors. In this study, we applied the Ensemble Algorithm for Clustering Cancer Data (EACCD) to create a prognostic system for lymphoma that integrates additional prognostic factors. Hodgkin and non-Hodgkin lymphoma survival data were extracted from the Surveillance, Epidemiology, and End Results Program of the National Cancer Institute and divided into the training set (131,725 cases) and the validation set (15,683 cases). Five prognostic factors were studied: Ann Arbor stage, type, site, age, and sex. EACCD was applied to the training set to produce a prognostic system, called an EACCD system, for convenience. The EACCD system stratified patients into eight prognostic groups with well-separated survival curves. These eight prognostic groups had significantly higher accuracies in survival prediction than the 24 Ann Arbor substages. A higher-risk group in the EACCD system roughly corresponds to a higher Ann Arbor substage. The proposed system shows a good performance in risk stratification and survival prediction on both the training and the validation sets. The EACCD system expands the traditional Ann Arbor staging system by leveraging additional prognostic information and is expected to advance treatment management for lymphoma patients.
Full article
(This article belongs to the Special Issue Feature Papers in Computational Biology and Medicine)
►▼
Show Figures
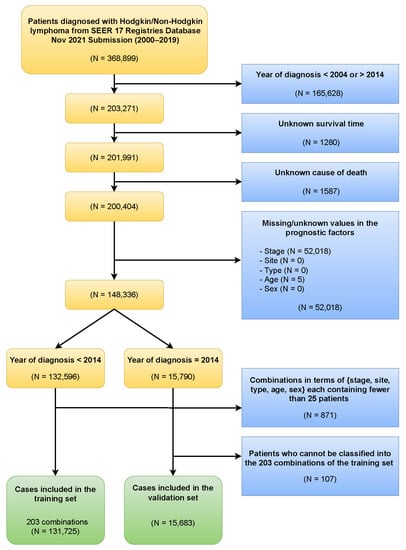
Figure 1
Open AccessArticle
Multimodal Deep Learning Methods on Image and Textual Data to Predict Radiotherapy Structure Names
by
, , , , , and
BioMedInformatics 2023, 3(3), 493-513; https://doi.org/10.3390/biomedinformatics3030034 - 25 Jun 2023
Abstract
Physicians often label anatomical structure sets in Digital Imaging and Communications in Medicine (DICOM) images with nonstandard random names. Hence, the standardization of these names for the Organs at Risk (OARs), Planning Target Volumes (PTVs), and ‘Other’ organs is a vital problem. This
[...] Read more.
Physicians often label anatomical structure sets in Digital Imaging and Communications in Medicine (DICOM) images with nonstandard random names. Hence, the standardization of these names for the Organs at Risk (OARs), Planning Target Volumes (PTVs), and ‘Other’ organs is a vital problem. This paper presents novel deep learning methods on structure sets by integrating multimodal data compiled from the radiotherapy centers of the US Veterans Health Administration (VHA) and Virginia Commonwealth University (VCU). These de-identified data comprise 16,290 prostate structures. Our method integrates the multimodal textual and imaging data with Convolutional Neural Network (CNN)-based deep learning approaches such as CNN, Visual Geometry Group (VGG) network, and Residual Network (ResNet) and shows improved results in prostate radiotherapy structure name standardization. Evaluation with macro-averaged F1 score shows that our model with single-modal textual data usually performs better than previous studies. The models perform well on textual data alone, while the addition of imaging data shows that deep neural networks achieve better performance using information present in other modalities. Additionally, using masked images and masked doses along with text leads to an overall performance improvement with the CNN-based architectures than using all the modalities together. Undersampling the majority class leads to further performance enhancement. The VGG network on the masked image-dose data combined with CNNs on the text data performs the best and presents the state-of-the-art in this domain.
Full article
(This article belongs to the Special Issue Advances in Quantitative Imaging Analysis: From Theory to Practice)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Detection of Myocardial Infarction Using Hybrid Models of Convolutional Neural Network and Recurrent Neural Network
BioMedInformatics 2023, 3(2), 478-492; https://doi.org/10.3390/biomedinformatics3020033 - 15 Jun 2023
Abstract
Myocardial Infarction (MI) is the death of the heart muscle caused by lack of oxygenated blood flow to the heart muscle. It has been the main cause of death worldwide. The fastest way to detect MI is by using an electrocardiogram (ECG) device,
[...] Read more.
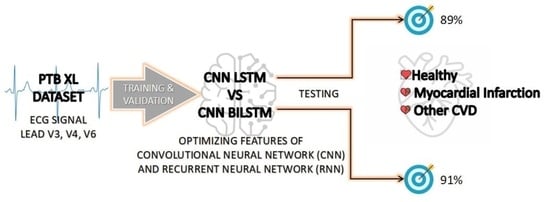
Myocardial Infarction (MI) is the death of the heart muscle caused by lack of oxygenated blood flow to the heart muscle. It has been the main cause of death worldwide. The fastest way to detect MI is by using an electrocardiogram (ECG) device, which generates graphs of heartbeats morphology over a certain period of time. Patients with MI need fast intervention as delay will lead to worsening heart conditions or failure. To improve MI diagnosis, much research has been carried out to come up with a fast and reliable system to aid automatic MI detection and prediction from ECG readings. Recurrent Neural Network (RNN) with memory has produced more accurate results in predicting time series problems. Convolutional neural networks have also shown good results in terms of solving prediction problems. However, CNN models do not have the capability of remembering temporal information. This research proposes hybrid models of CNN and RNN techniques to predict MI. Specifically, CNN-LSTM and CNN-BILSTM models have been developed. The PTB XL dataset is used to train the models. The models predict ECG input as representing MI symptoms, healthy heart conditions or other cardiovascular diseases. Deep learning models offer automatic feature extraction, and our models take advantage of automatic feature extraction. The other superior models used their own feature extraction algorithm. This research proposed a straightforward architecture that depends mostly on the capability of the deep learning model to learn the data. Performance evaluation of the models shows overall accuracy of 89% for CNN LSTM and 91% for the CNN BILSTM model.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Uncovering Disease-Related Polymorphisms through Correlations between SNP Frequencies, Population and Epidemiological Data
BioMedInformatics 2023, 3(2), 467-477; https://doi.org/10.3390/biomedinformatics3020032 - 13 Jun 2023
Abstract
Background: According to GWAS, which analyzes large amounts of DNA variants in case-control strategies, the genetic differences between two human individuals do not exceed 0.5%. As a consequence, finding biological significance in GWAS results is a challenging task. We propose an alternative method
[...] Read more.
Background: According to GWAS, which analyzes large amounts of DNA variants in case-control strategies, the genetic differences between two human individuals do not exceed 0.5%. As a consequence, finding biological significance in GWAS results is a challenging task. We propose an alternative method for identifying disease-causing variants based on the simultaneous evaluation of genome variant data acquired from public databases and pathology epidemiological data. This method is grounded on the following premise: If a particular pathology is common in a community, genetic variants that confer susceptibility to that pathology should also be common in that population. Methods: Three groups of genes were evaluated to test this premise: variants related to depression found through GWAS, six genes unrelated to depression, and four genes already genotyped in case-control studies involving depression (TPH2, NR3C1, SLC6A2 and SLC6A3). In terms of GWAS depression-related variants, nine of the 82 SNPs evaluated showed a favorable correlation between allele frequency and epidemiological data. As anticipated, none of the 286 SNPs were correlated in the neutral group. In terms of proof of concept, two THP2 variants, 26 NR3C1 variants and four SLC6A3 variants were found to be related to depression rates and epidemiological statistics. Conclusions: Together with data from the literature involving these SNPs, these correlations support this strategy as a complementary method for identifying possible disease-causing variants.
Full article
(This article belongs to the Section Applied Biomedical Data Science)
►▼
Show Figures
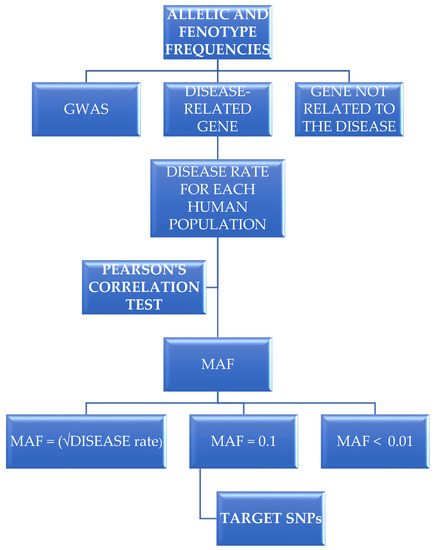
Figure 1
Open AccessArticle
Automatic Facial Palsy, Age and Gender Detection Using a Raspberry Pi
BioMedInformatics 2023, 3(2), 455-466; https://doi.org/10.3390/biomedinformatics3020031 - 13 Jun 2023
Abstract
Facial palsy (FP) is a neurological disorder that affects the facial nerve, specifically the seventh nerve, resulting in the patient losing control of the facial muscles on one side of the face. It is an annoying condition that can occur in both children
[...] Read more.
Facial palsy (FP) is a neurological disorder that affects the facial nerve, specifically the seventh nerve, resulting in the patient losing control of the facial muscles on one side of the face. It is an annoying condition that can occur in both children and adults, regardless of gender. Diagnosis by visual examination, based on differences in the sides of the face, can be prone to errors and inaccuracies. The detection of FP using artificial intelligence through computer vision systems has become increasingly important. Deep learning is the best solution for detecting FP in real-time with high accuracy, saving patients time, effort, and cost. Therefore, this work proposes a real-time detection system for FP, and for determining the patient’s gender and age, using a Raspberry Pi device with a digital camera and a deep learning algorithm. The solution facilitates the diagnosis process for both the doctor and the patient, and it could be part of a medical assessment activity. This study used a dataset of 20,600 images, containing 19,000 normal images and 1600 FP images, to achieve an accuracy of 98%. Thus, the proposed system is a highly accurate and capable medical diagnostic tool for detecting FP.
Full article
(This article belongs to the Special Issue Deep Learning Methods and Application for Bioinformatics and Healthcare)
►▼
Show Figures
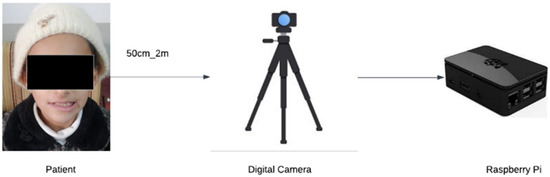
Figure 1
Open AccessArticle
An IoT-Based Automatic and Continuous Urine Measurement System
BioMedInformatics 2023, 3(2), 446-454; https://doi.org/10.3390/biomedinformatics3020030 - 05 Jun 2023
Abstract
Urine output is an important indicator of renal function. In hospitals, urine is collected using a catheter connected to a urine collection bag that has volume gradation markings. This type of visual measurement has low levels of accuracy and is labor-intensive. This paper
[...] Read more.
Urine output is an important indicator of renal function. In hospitals, urine is collected using a catheter connected to a urine collection bag that has volume gradation markings. This type of visual measurement has low levels of accuracy and is labor-intensive. This paper developed an Internet-of-Things enabled system that continuously monitors the urine volume collected via the urine collection system. The device is built utilizing a strain gauge load cell, an integrated circuit that contains an amplifier, analog-to-digital converter, and a WiFi-enabled microcontroller. The data is sent via wireless networking to a data collection and analysis server, which provides accurate analyses of urine output. A mobile application utilizing the Blynk.io system is used to display the data. This device and mobile application were built at a minimal cost of 26 USD. The device has been tested multiple times and reported urine output accurately, with minimal difference between actual versus measured volumes. In the future, further development of this device can provide hospitals and physicians worldwide with easy access to affordable, accurate, and real-time urine measurement, which would translate into better, life-saving medical care.
Full article
(This article belongs to the Section Computational Biology and Medicine)
►▼
Show Figures
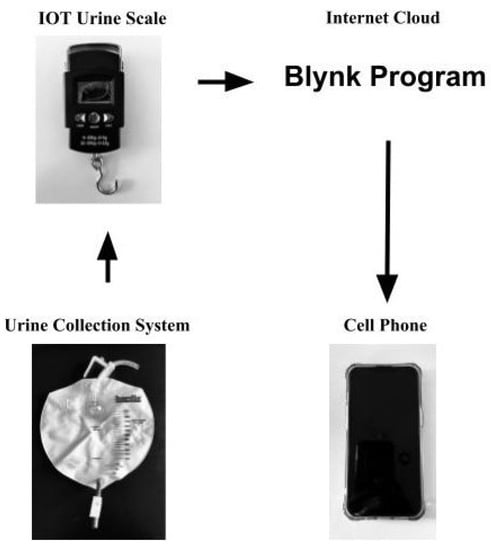
Figure 1
Open AccessArticle
Integrative Molecular Analysis of DNA Methylation Dynamics Unveils Molecules with Prognostic Potential in Breast Cancer
by
, , , , , , , and
BioMedInformatics 2023, 3(2), 434-445; https://doi.org/10.3390/biomedinformatics3020029 - 05 Jun 2023
Abstract
DNA methylation acts as a major epigenetic modification in mammals, characterized by the transfer of a methyl group to a cytosine. DNA methylation plays a pivotal role in regulating normal development, and misregulation in cells leads to an abnormal phenotype as is seen
[...] Read more.
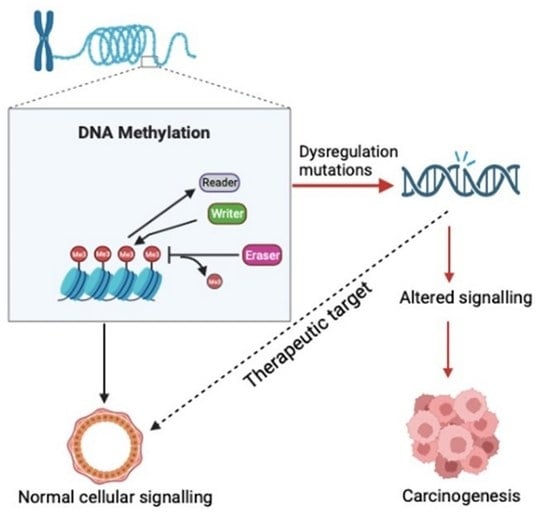
DNA methylation acts as a major epigenetic modification in mammals, characterized by the transfer of a methyl group to a cytosine. DNA methylation plays a pivotal role in regulating normal development, and misregulation in cells leads to an abnormal phenotype as is seen in several cancers. Any mutations or expression anomalies of genes encoding regulators of DNA methylation may lead to abnormal expression of critical molecules. A comprehensive genomic study encompassing all the genes related to DNA methylation regulation in relation to breast cancer is lacking. We used genomic and transcriptomic datasets from the Cancer Genome Atlas (TGCA) Pan-Cancer Atlas, Genotype-Tissue Expression (GTEx) and microarray platforms and conducted in silico analysis of all the genes related to DNA methylation with respect to writing, reading and erasing this epigenetic mark. Analysis of mutations was conducted using cBioportal, while Xena and KMPlot were utilized for expression changes and patient survival, respectively. Our study identified multiple mutations in the genes encoding regulators of DNA methylation. The expression profiling of these showed significant differences between normal and disease tissues. Moreover, deregulated expression of some of the genes, namely DNMT3B, MBD1, MBD6, BAZ2B, ZBTB38, KLF4, TET2 and TDG, was correlated with patient prognosis. The current study, to our best knowledge, is the first to provide a comprehensive molecular and genetic profile of DNA methylation machinery genes in breast cancer and identifies DNA methylation machinery as an important determinant of the disease progression. The findings of this study will advance our understanding of the etiology of the disease and may serve to identify alternative targets for novel therapeutic strategies in cancer.
Full article
(This article belongs to the Special Issue Feature Papers in Computational Biology and Medicine)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Reinforcement Learning for Multiple Daily Injection (MDI) Therapy in Type 1 Diabetes (T1D)
by
and
BioMedInformatics 2023, 3(2), 422-433; https://doi.org/10.3390/biomedinformatics3020028 - 05 Jun 2023
Abstract
In this study, we propose a closed-loop insulin administration framework for multiple daily injection (MDI) treatment using a reinforcement learning (RL) agent for insulin bolus therapy. The RL agent, based on the soft actor–critic (SAC) algorithm, dynamically adjusts insulin dosages based on real-time
[...] Read more.
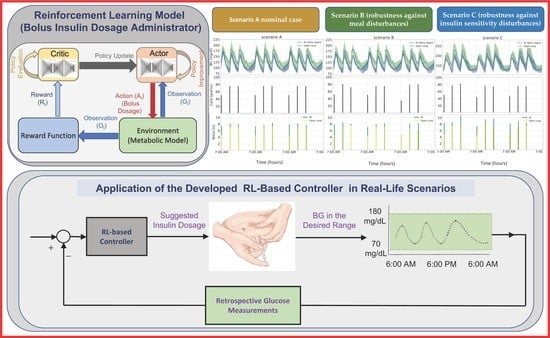
In this study, we propose a closed-loop insulin administration framework for multiple daily injection (MDI) treatment using a reinforcement learning (RL) agent for insulin bolus therapy. The RL agent, based on the soft actor–critic (SAC) algorithm, dynamically adjusts insulin dosages based on real-time glucose readings, meal intakes, and previous actions. We evaluated the proposed strategy on ten in silico patients with type 1 diabetes undergoing MDI therapy, considering three meal scenarios. The results show that, compared to an open-loop conventional therapy, our proposed closed-loop control strategy significantly reduces glucose variability and increases the percentage of time the glucose levels remained within the target range. In particular, the weekly mean glucose level reduced from 145.34 ± 57.26 mg/dL to 115.18 ± 7.93 mg/dL, 143.62 ± 55.72 mg/dL to 115.28 ± 8.11 mg/dL, and 171.63 ± 49.30 mg/dL to 143.94 ± 23.81 mg/dL for Scenarios A, B and C, respectively. Furthermore, the percent time in range (70–180 mg/dL) significantly improved from 63.77 ± 27.90% to 91.72 ± 9.27% (p = 0.01) in Scenario A, 64.82 ± 28.06% to 92.29 ± 9.15% (p = 0.01) in Scenario B, and 58.45 ± 27.53% to 81.45 ± 26.40% (p = 0.05) in Scenario C. The model also demonstrated robustness against meal disturbances and insulin sensitivity disturbances, achieving mean glucose levels within the target range and maintaining a low risk of hypoglycemia, which were statistically significant for Scenarios B and C. The proposed model outperformed open-loop conventional therapy in all scenarios, highlighting the potential of RL-based closed-loop insulin administration models in improving diabetes management.
Full article
(This article belongs to the Section Clinical Informatics)
►▼
Show Figures
Graphical abstract
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
AI, BioMedInformatics, Cancers, Current Oncology, Medical Sciences
Uro-Oncology in the Robotic Era: Advancements, Limitations, and Further Perspectives
Topic Editors: Cosimo De Nunzio, Aldo BrassettiDeadline: 1 September 2023
Topic in
BioMedInformatics, Cancers, Current Oncology, Diagnostics, Healthcare
Real-Time Monitoring for Improving Cancer Diagnosis and Prognosis
Topic Editors: Maria Li Lung, Josephine KoDeadline: 19 October 2023
Topic in
Applied Sciences, BioMedInformatics, BioTech, Genes, Computation
Computational Intelligence and Bioinformatics (CIB)
Topic Editors: Marco Mesiti, Giorgio Valentini, Elena Casiraghi, Tiffany J. CallahanDeadline: 31 October 2023
Topic in
BioMed, Biomedicines, BioMedInformatics, Epidemiologia, JCM
Health Informatics and Epidemiological Data Analysis in COVID-19 Based Internet of Medical Things (HIEDA-COVID19-IOMT)
Topic Editors: Wenjun (Chris) Zhang, Dhanjoo N. Ghista, Kelvin K.L. Wong, Zhili SunDeadline: 15 December 2023

Conferences
Special Issues
Special Issue in
BioMedInformatics
Advances in Quantitative Imaging Analysis: From Theory to Practice
Guest Editors: Federico Mastroleo, Angela Ammirabile, Giulia MarvasoDeadline: 30 November 2023
Special Issue in
BioMedInformatics
Application of Semantic Web Technologies in Biomedicine and Biomedical Informatics
Guest Editors: Pietro Hiram Guzzi, Pietro CinagliaDeadline: 28 December 2023
Special Issue in
BioMedInformatics
Feature Papers in Medical Statistics and Data Science Section
Guest Editor: Pentti NieminenDeadline: 31 December 2023
Special Issue in
BioMedInformatics
Feature Papers in Computational Biology and Medicine
Guest Editor: Hans BinderDeadline: 31 July 2024