Journal Description
Epigenomes
Epigenomes
is an international, peer-reviewed, open access journal on epigenetics and epigenomics, published quarterly online by MDPI. The Epigenetics Society is affiliated with Epigenomes and its members receive discounts on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, ESCI (Web of Science), PMC, PubMed, Embase, PubAg, CAPlus / SciFinder, and other databases.
- Journal Rank: CiteScore - Q2 (Biochemistry, Genetics and Molecular Biology (miscellaneous))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 22.6 days after submission; acceptance to publication is undertaken in 4.6 days (median values for papers published in this journal in the first half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
Impact Factor:
2.5 (2022);
5-Year Impact Factor:
2.1 (2022)
Latest Articles
Multiomics Data Analysis Identified CpG Sites That Mediate the Impact of Smoking on Cardiometabolic Traits
Epigenomes 2023, 7(3), 19; https://doi.org/10.3390/epigenomes7030019 - 22 Aug 2023
Abstract
►
Show Figures
Understanding the epigenome paths through which smoking contributes to cardiometabolic traits is important for downstream applications. In this study, an SNP-based analytical pipeline was used to integrate several publicly available datasets in order to identify CpG sites that mediate the impact of smoking
[...] Read more.
Understanding the epigenome paths through which smoking contributes to cardiometabolic traits is important for downstream applications. In this study, an SNP-based analytical pipeline was used to integrate several publicly available datasets in order to identify CpG sites that mediate the impact of smoking on cardiometabolic traits and to investigate the underlying molecular mechanisms. After applying stringent statistical criteria, 11 CpG sites were detected that showed significant association (p < 5 × 10−8) with cardiometabolic traits at both the discovery and replication stages. By integrating eQTL data, I found genes behind a number of these associations. cg05228408 was hypomethylated in smokers and contributed to higher blood pressure by lowering the expression of the CLCN6 gene. cg08639339 was hypermethylated in smokers and lowered the metabolic rate by increasing the expression of RAB29; furthermore, I noted TMEM120A mediated the impact of smoking-cg17325771 on LDL, and LTBP3 mediated the smoking-cg07029024 effect on heart rate. The pathway analysis identified processes through which the identified genes impact their traits. This study provides a list of CpG sites that mediates the impact of smoking on cardiometabolic traits and a framework to investigate the underlying molecular paths using publicly available data.
Full article
Open AccessArticle
Excessive Gestational Weight Gain Alters DNA Methylation and Influences Foetal and Neonatal Body Composition
by
, , , , and
João Victor da Silva Guerra
Epigenomes 2023, 7(3), 18; https://doi.org/10.3390/epigenomes7030018 - 16 Aug 2023
Abstract
►▼
Show Figures
Background: Changes in body weight are associated with the regulation of DNA methylation (DNAm). In this study, we investigated the associations between maternal gestational weight gain-related DNAm and foetal and neonatal body composition. Methods: Brazilian pregnant women from the Araraquara Cohort Study were
[...] Read more.
Background: Changes in body weight are associated with the regulation of DNA methylation (DNAm). In this study, we investigated the associations between maternal gestational weight gain-related DNAm and foetal and neonatal body composition. Methods: Brazilian pregnant women from the Araraquara Cohort Study were followed up during pregnancy, delivery, and after hospital discharge. Women with normal pre-pregnancy BMI were allocated into two groups: adequate gestational weight gain (AGWG, n = 45) and excessive gestational weight gain (EGWG, n = 30). Foetal and neonatal body composition was evaluated via ultrasound and plethysmography, respectively. DNAm was assessed in maternal blood using Illumina Infinium MethylationEPIC BeadChip arrays. Linear regression models were used to explore the associations between DNAm and foetal and neonatal body composition. Results: Maternal weight, GWG, neonatal weight, and fat mass were higher in the EGWG group. Analysis of DNAm identified 46 differentially methylated positions and 11 differentially methylated regions (DMRs) between the EGWG and AGWG groups. Nine human phenotypes were enriched for these 11 DMRs located in 13 genes (EMILIN1, HOXA5, CPT1B, CLDN9, ZFP57, BRCA1, POU5F1, ANKRD33, HLA-B, RANBP17, ZMYND11, DIP2C, TMEM232), highlighting the terms insulin resistance, and hyperglycaemia. Maternal DNAm was associated with foetal total thigh and arm tissues and subcutaneous thigh and arm fat, as well as with neonatal fat mass percentage and fat mass. Conclusion: The methylation pattern in the EGWG group indicated a risk for developing chronic diseases and involvement of maternal DNAm in foetal lean and fat mass and in neonatal fat mass.
Full article
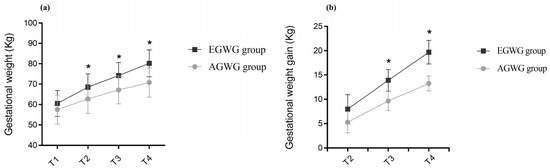
Figure 1
Open AccessReview
Ribosomal Biogenesis and Heterogeneity in Development, Disease, and Aging
Epigenomes 2023, 7(3), 17; https://doi.org/10.3390/epigenomes7030017 - 11 Aug 2023
Abstract
►▼
Show Figures
Although reported in the literature, ribosome heterogeneity is a phenomenon whose extent and implications in cell and organismal biology is not fully appreciated. This has been the case due to the lack of the appropriate techniques and approaches. Heterogeneity can arise from alternative
[...] Read more.
Although reported in the literature, ribosome heterogeneity is a phenomenon whose extent and implications in cell and organismal biology is not fully appreciated. This has been the case due to the lack of the appropriate techniques and approaches. Heterogeneity can arise from alternative use and differential content of protein and RNA constituents, as well as from post-transcriptional and post-translational modifications. In the few examples we have, it is apparent that ribosomal heterogeneity offers an additional level and potential for gene expression regulation and might be a way towards tuning metabolism, stress, and growth programs to external and internal stimuli and needs. Here, we introduce ribosome biogenesis and discuss ribosomal heterogeneity in various reported occasions. We conclude that a systematic approach in multiple organisms will be needed to delineate this biological phenomenon and its contributions to growth, aging, and disease. Finally, we discuss ribosome mutations and their roles in disease.
Full article
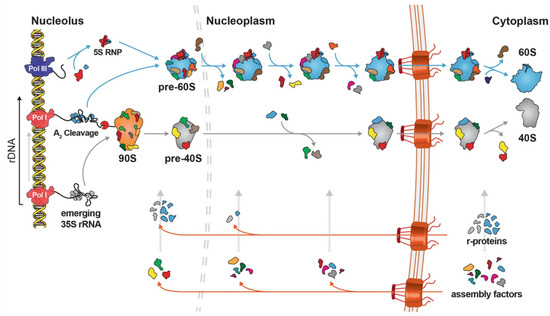
Figure 1
Open AccessReview
Evaluation of Exosomal miRNA as Potential Biomarkers in Cervical Cancer
by
, , , , , , and
Epigenomes 2023, 7(3), 16; https://doi.org/10.3390/epigenomes7030016 - 01 Aug 2023
Abstract
Different studies show that small non-coding RNAs, such as microRNAs (miRNAs) obtained from exosomes, are considered potential biomarkers in several types of cancer, including cervical cancer (CC). Therefore, the present study seeks to present an overview of the role of circulating exosomal miRNAs
[...] Read more.
Different studies show that small non-coding RNAs, such as microRNAs (miRNAs) obtained from exosomes, are considered potential biomarkers in several types of cancer, including cervical cancer (CC). Therefore, the present study seeks to present an overview of the role of circulating exosomal miRNAs with the potential to act as biomarkers for the diagnosis and prognosis of CC and to analyze the presence of these miRNAs according to the stage of CC. For this purpose, a review was developed, with articles consulted from the electronic databases MEDLINE/PubMed, Scopus, and Web of Science published between 2015 and 2021. Seven articles were included after a selection of studies according to the eligibility criteria. In addition to the methods used for sample analysis, detection, and isolation of miRNAs in each article, clinical data were also extracted from the patients studied, such as the stage of cancer. After analyzing the network of the seven miRNAs, they were associated with the immune system, CC progression and staging, and cisplatin resistance. With the belief that studies on miRNAs in cervical cancer would have major clinical implications, in this review, we have attempted to summarize the current situation and potential development prospects.
Full article
(This article belongs to the Special Issue Epigenetic Regulation in Cancer)
►▼
Show Figures

Figure 1
Open AccessArticle
Association of Toll-Like Receptor Gene Polymorphisms with Tuberculosis in HIV-Positive Participants
by
, , , , , and
Epigenomes 2023, 7(3), 15; https://doi.org/10.3390/epigenomes7030015 - 25 Jul 2023
Abstract
Genetic factors in the HIV-background may play a significant role in the susceptibility to secondary diseases, like tuberculosis, which is the leading cause in mortality of HIV-positive people. Toll-like receptors (TLRs) are considered to be receptors for adaptive immunity, and polymorphisms in TLR
[...] Read more.
Genetic factors in the HIV-background may play a significant role in the susceptibility to secondary diseases, like tuberculosis, which is the leading cause in mortality of HIV-positive people. Toll-like receptors (TLRs) are considered to be receptors for adaptive immunity, and polymorphisms in TLR genes can influence the activity of the immune response to infection. We conducted a case–control study of the association of TLR gene polymorphisms with the risk of tuberculosis coinfection in a multi-country sample of HIV-positive participants. Our study revealed certain associations between TLR4 and TLR6 polymorphisms and HIV–tuberculosis coinfection. We also found that the analyzed TLR1 and TLR4 polymorphisms were linked with the decline in CD4+ cell count, which is a predictor of disease progression in HIV-infected individuals. Our findings confirm that TLR gene polymorphisms are factors that may contribute to development of HIV–tuberculosis coinfection. However, the essence of the observed associations remains unclear, since it can also include both environmental factors and epigenetic mechanisms of gene expression regulation.
Full article
(This article belongs to the Topic Biosocial Studies)
►▼
Show Figures
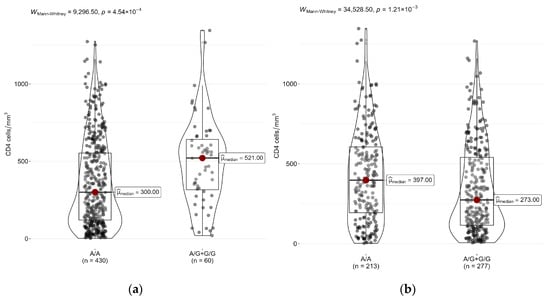
Figure 1
Open AccessReview
Epigenetic Regulation in Heterosis and Environmental Stress: The Challenge of Producing Hybrid Epigenomes to Face Climate Change
Epigenomes 2023, 7(3), 14; https://doi.org/10.3390/epigenomes7030014 - 24 Jul 2023
Abstract
Epigenetic regulation has the potential to revolutionize plant breeding and improve crop yields by regulating gene expression in plants. DNA methylation and histone modifications are key epigenetic modifications that can impact plant development, stress responses, productivity, and yields. Higher-yielding crops not only generate
[...] Read more.
Epigenetic regulation has the potential to revolutionize plant breeding and improve crop yields by regulating gene expression in plants. DNA methylation and histone modifications are key epigenetic modifications that can impact plant development, stress responses, productivity, and yields. Higher-yielding crops not only generate greater profits for farmers and seed producers, but also require less land, water, fuel, and fertilizer than traditional crops for equivalent yields. The use of heterosis in crops can influence productivity and food quality, but producing hybrids with superior agronomic traits to their parents remains challenging. However, epigenetic markers, such as histone methylation and acetylation, may help select parental and hybrid combinations with better performances than the parental plants. This review assesses the potential applications of epigenetics in crop breeding and improvement, rendering agriculture more efficient, sustainable, and adaptable to changing environmental conditions.
Full article
(This article belongs to the Special Issue Plant Epigenome and Epitranscriptome Adaptation to Environmental Changes)
►▼
Show Figures
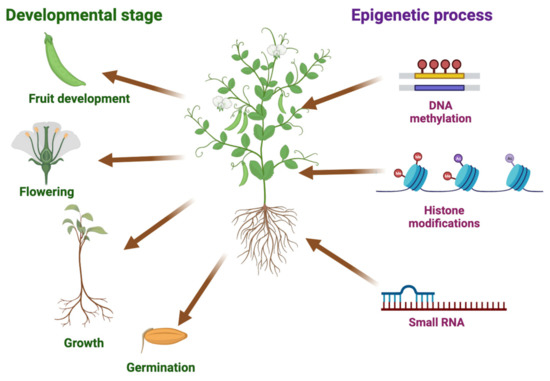
Figure 1
Open AccessReview
Pulmonary Pathogen-Induced Epigenetic Modifications
Epigenomes 2023, 7(3), 13; https://doi.org/10.3390/epigenomes7030013 - 06 Jul 2023
Abstract
►▼
Show Figures
Epigenetics generally involves genetic control by factors other than our own DNA sequence. Recent research has focused on delineating the mechanisms of two major epigenetic phenomena: DNA methylation and histone modification. As epigenetics involves many cellular processes, it is no surprise that it
[...] Read more.
Epigenetics generally involves genetic control by factors other than our own DNA sequence. Recent research has focused on delineating the mechanisms of two major epigenetic phenomena: DNA methylation and histone modification. As epigenetics involves many cellular processes, it is no surprise that it can also influence disease-associated gene expression. A direct link between respiratory infections, host cell epigenetic regulations, and chronic lung diseases is still unknown. Recent studies have revealed bacterium- or virus-induced epigenetic changes in the host cells. In this review, we focused on respiratory pathogens (viruses, bacteria, and fungi) induced epigenetic modulations (DNA methylation and histone modification) that may contribute to lung disease pathophysiology by promoting host defense or allowing pathogen persistence.
Full article

Figure 1
Open AccessReview
Therapeutic Applications of Azanucleoside Analogs as DNA Demethylating Agents
by
and
Epigenomes 2023, 7(3), 12; https://doi.org/10.3390/epigenomes7030012 - 05 Jul 2023
Abstract
►▼
Show Figures
Azanucleosides, such as 5-azacytidine and decitabine, are DNA demethylating agents used in the treatment of acute myeloid leukemia and myelodysplastic syndromes. Researchers continue to explore their utility in the treatment of other hematologic and solid tumors. Based on the capacity of the compounds
[...] Read more.
Azanucleosides, such as 5-azacytidine and decitabine, are DNA demethylating agents used in the treatment of acute myeloid leukemia and myelodysplastic syndromes. Researchers continue to explore their utility in the treatment of other hematologic and solid tumors. Based on the capacity of the compounds to inhibit DNA methyltransferase enzymes and the important role of DNA methylation in health and disease, it is essential to understand the molecular changes that azanucleosides induce and how these changes may improve treatment outcomes in subsets of patients. This review summarizes the molecular and therapeutic actions of azanucleosides and discusses recent clinical trials of these compounds as single agents or in combination therapy for the treatment of cancer and related conditions.
Full article

Figure 1
Open AccessArticle
Graft-Derived Cell-Free DNA Quantification following Liver Transplantation Using Tissue-Specific DNA Methylation and Donor-Specific Genotyping Techniques: An Orthogonal Comparison Study
by
, , , , , , and
Epigenomes 2023, 7(2), 11; https://doi.org/10.3390/epigenomes7020011 - 09 Jun 2023
Abstract
►▼
Show Figures
Background: Graft-derived cell-free DNA (gdcfDNA) analysis has shown promise as a non-invasive tool for monitoring organ health following solid organ transplantation. A number of gdcfDNA analysis techniques have been described; however, the majority rely on sequencing or prior genotyping to detect donor-recipient
[...] Read more.
Background: Graft-derived cell-free DNA (gdcfDNA) analysis has shown promise as a non-invasive tool for monitoring organ health following solid organ transplantation. A number of gdcfDNA analysis techniques have been described; however, the majority rely on sequencing or prior genotyping to detect donor-recipient mis-matched genetic polymorphisms. Differentially methylated regions of DNA can be used to identify the tissue-of-origin of cell-free DNA (cfDNA) fragments. In this study, we aimed to directly compare the performance of gdcfDNA monitoring using graft-specific DNA methylation analysis and donor-recipient genotyping techniques in a pilot cohort of clinical samples from patients post-liver transplantation. Results: 7 patients were recruited prior to LT, 3 developed early, biopsy-proven TCMR in the first 6 weeks post-LT. gdcfDNA was successfully quantified in all samples using both approaches. There was a high level of technical correlation between results using the two techniques (Spearman testing, rs = 0.87, p < 0.0001). gdcfDNA levels quantified using the genotyping approach were significantly greater across all timepoints in comparison to the tissue-specific DNA methylation-based approach: e.g., day 1 post-LT median 31,350 copies/mL (IQR 6731–64,058) vs. 4133 copies/mL (IQR 1100–8422), respectively. Qualitative trends in gdcfDNA levels for each patient were concordant between the two assays. Acute TCMR was preceded by significant elevations in gdcfDNA as quantified by both techniques. Elevations in gdcfDNA, using both techniques, were suggestive of TCMR in this pilot study with a 6- and 3-day lead-time prior to histological diagnosis in patients 1 and 2. Conclusions: Both the graft-specific methylation and genotyping techniques successfully quantified gdcfDNA in patients post-LT with statistically significant concordance. A direct comparison of these two techniques is not only important from a technical perspective for orthogonal validation, but significantly adds weight to the evidence that gdcfDNA monitoring reflects the underlying biology. Both techniques identified LT recipients who developed acute TCMR, with several days lead-time in comparison to conventional diagnostic workflows. Whilst the two assays performed comparably, gdcfDNA monitoring based on graft-specific DNA methylation patterns in cfDNA offers major practical advantages over the donor-recipient genotyping, and hence enhances the potential to translate this emerging technology into clinical practice.
Full article
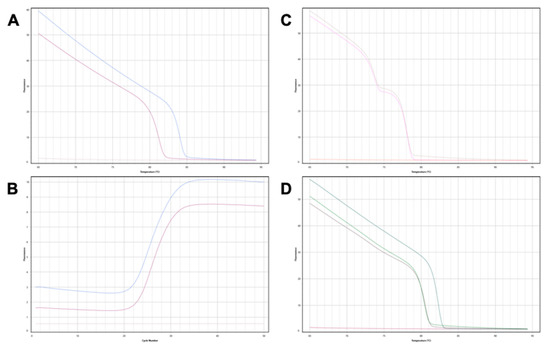
Figure 1
Open AccessReview
Histone Demethylase Modulation: Epigenetic Strategy to Combat Cancer Progression
Epigenomes 2023, 7(2), 10; https://doi.org/10.3390/epigenomes7020010 - 17 May 2023
Abstract
Epigenetic modifications are heritable, reversible changes in histones or the DNA that control gene functions, being exogenous to the genomic sequence itself. Human diseases, particularly cancer, are frequently connected to epigenetic dysregulations. One of them is histone methylation, which is a dynamically reversible
[...] Read more.
Epigenetic modifications are heritable, reversible changes in histones or the DNA that control gene functions, being exogenous to the genomic sequence itself. Human diseases, particularly cancer, are frequently connected to epigenetic dysregulations. One of them is histone methylation, which is a dynamically reversible and synchronously regulated process that orchestrates the three-dimensional epigenome, nuclear processes of transcription, DNA repair, cell cycle, and epigenetic functions, by adding or removing methylation groups to histones. Over the past few years, reversible histone methylation has become recognized as a crucial regulatory mechanism for the epigenome. With the development of numerous medications that target epigenetic regulators, epigenome-targeted therapy has been used in the treatment of malignancies and has shown meaningful therapeutic potential in preclinical and clinical trials. The present review focuses on the recent advances in our knowledge on the role of histone demethylases in tumor development and modulation, in emphasizing molecular mechanisms that control cancer cell progression. Finally, we emphasize current developments in the advent of new molecular inhibitors that target histone demethylases to regulate cancer progression.
Full article
(This article belongs to the Special Issue Recent Advances in Biological Methylation 2022)
►▼
Show Figures
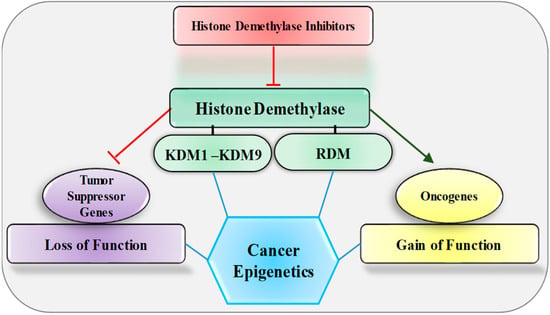
Figure 1
Open AccessReview
Critical Considerations for Investigating MicroRNAs during Tumorigenesis: A Case Study in Conceptual and Contextual Nuances of miR-211-5p in Melanoma
Epigenomes 2023, 7(2), 9; https://doi.org/10.3390/epigenomes7020009 - 26 Apr 2023
Abstract
MicroRNAs are non-coding RNAs fundamental to metazoan development and disease. Although the aberrant regulation of microRNAs during mammalian tumorigenesis is well established, investigations into the contributions of individual microRNAs are wrought with conflicting observations. The underlying cause of these inconsistencies is often attributed
[...] Read more.
MicroRNAs are non-coding RNAs fundamental to metazoan development and disease. Although the aberrant regulation of microRNAs during mammalian tumorigenesis is well established, investigations into the contributions of individual microRNAs are wrought with conflicting observations. The underlying cause of these inconsistencies is often attributed to context-specific functions of microRNAs. We propose that consideration of both context-specific factors, as well as underappreciated fundamental concepts of microRNA biology, will permit a more harmonious interpretation of ostensibly diverging data. We discuss the theory that the biological function of microRNAs is to confer robustness to specific cell states. Through this lens, we then consider the role of miR-211-5p in melanoma progression. Using literature review and meta-analyses, we demonstrate how a deep understating of domain-specific contexts is critical for moving toward a concordant understanding of miR-211-5p and other microRNAs in cancer biology.
Full article
(This article belongs to the Collection Epigenetics of Melanoma)
►▼
Show Figures
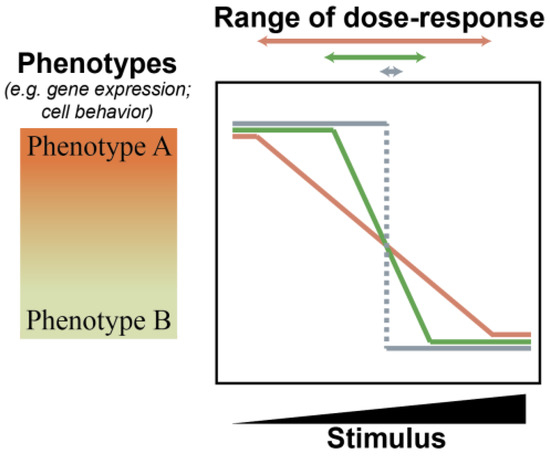
Figure 1
Open AccessArticle
Epigenome-Wide Changes in the Cell Layers of the Vein Wall When Exposing the Venous Endothelium to Oscillatory Shear Stress
by
, , , , , and
Epigenomes 2023, 7(1), 8; https://doi.org/10.3390/epigenomes7010008 - 20 Mar 2023
Abstract
►▼
Show Figures
Epigenomic changes in the venous cells exerted by oscillatory shear stress towards the endothelium may result in consolidation of gene expression alterations upon vein wall remodeling during varicose transformation. We aimed to reveal such epigenome-wide methylation changes. Primary culture cells were obtained from
[...] Read more.
Epigenomic changes in the venous cells exerted by oscillatory shear stress towards the endothelium may result in consolidation of gene expression alterations upon vein wall remodeling during varicose transformation. We aimed to reveal such epigenome-wide methylation changes. Primary culture cells were obtained from non-varicose vein segments left after surgery of 3 patients by growing the cells in selective media after magnetic immunosorting. Endothelial cells were either exposed to oscillatory shear stress or left at the static condition. Then, other cell types were treated with preconditioned media from the adjacent layer’s cells. DNA isolated from the harvested cells was subjected to epigenome-wide study using Illumina microarrays followed by data analysis with GenomeStudio (Illumina), Excel (Microsoft), and Genome Enhancer (geneXplain) software packages. Differential (hypo-/hyper-) methylation was revealed for each cell layer’s DNA. The most targetable master regulators controlling the activity of certain transcription factors regulating the genes near the differentially methylated sites appeared to be the following: (1) HGS, PDGFB, and AR for endothelial cells; (2) HGS, CDH2, SPRY2, SMAD2, ZFYVE9, and P2RY1 for smooth muscle cells; and (3) WWOX, F8, IGF2R, NFKB1, RELA, SOCS1, and FXN for fibroblasts. Some of the identified master regulators may serve as promising druggable targets for treating varicose veins in the future.
Full article
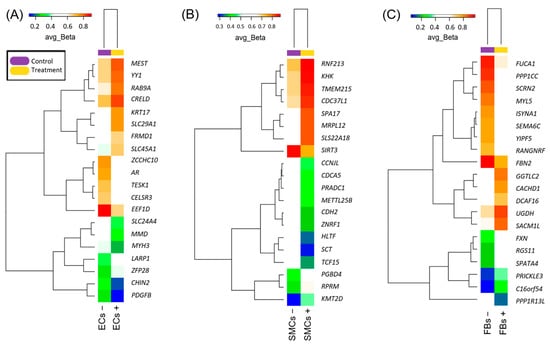
Figure 1
Open AccessReview
Chemical Inhibitors Targeting the Histone Lysine Demethylase Families with Potential for Drug Discovery
Epigenomes 2023, 7(1), 7; https://doi.org/10.3390/epigenomes7010007 - 11 Mar 2023
Abstract
The dynamic regulation of histone methylation and demethylation plays an important role in the regulation of gene expression. Aberrant expression of histone lysine demethylases has been implicated in various diseases including intractable cancers, and thus lysine demethylases serve as promising therapeutic targets. Recent
[...] Read more.
The dynamic regulation of histone methylation and demethylation plays an important role in the regulation of gene expression. Aberrant expression of histone lysine demethylases has been implicated in various diseases including intractable cancers, and thus lysine demethylases serve as promising therapeutic targets. Recent studies in epigenomics and chemical biology have led to the development of a series of small-molecule demethylase inhibitors that are potent, specific, and have in vivo efficacy. In this review, we highlight emerging small-molecule inhibitors targeting the histone lysine demethylases and their progress toward drug discovery.
Full article
(This article belongs to the Special Issue Epidrugs: Toward Understanding and Treating Diverse Diseases)
►▼
Show Figures
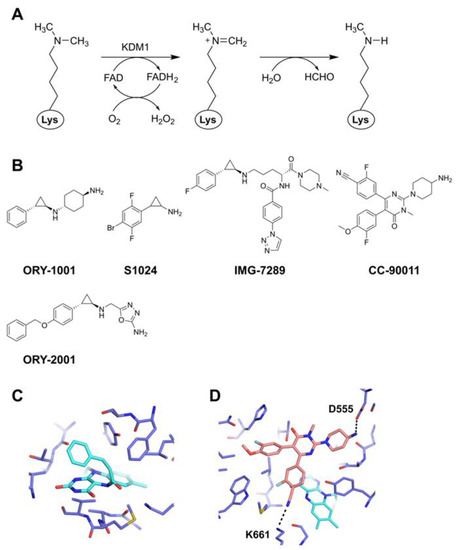
Figure 1
Open AccessReview
Epigenetic Regulation in Breast Cancer: Insights on Epidrugs
Epigenomes 2023, 7(1), 6; https://doi.org/10.3390/epigenomes7010006 - 18 Feb 2023
Cited by 4
Abstract
Breast cancer remains a common cause of cancer-related death in women. Therefore, further studies are necessary for the comprehension of breast cancer and the revolution of breast cancer treatment. Cancer is a heterogeneous disease that results from epigenetic alterations in normal cells. Aberrant
[...] Read more.
Breast cancer remains a common cause of cancer-related death in women. Therefore, further studies are necessary for the comprehension of breast cancer and the revolution of breast cancer treatment. Cancer is a heterogeneous disease that results from epigenetic alterations in normal cells. Aberrant epigenetic regulation is strongly associated with the development of breast cancer. Current therapeutic approaches target epigenetic alterations rather than genetic mutations due to their reversibility. The formation and maintenance of epigenetic changes depend on specific enzymes, including DNA methyltransferases and histone deacetylases, which are promising targets for epigenetic-based therapy. Epidrugs target different epigenetic alterations, including DNA methylation, histone acetylation, and histone methylation, which can restore normal cellular memory in cancerous diseases. Epigenetic-targeted therapy using epidrugs has anti-tumor effects on malignancies, including breast cancer. This review focuses on the importance of epigenetic regulation and the clinical implications of epidrugs in breast cancer.
Full article
(This article belongs to the Special Issue Epidrugs: Toward Understanding and Treating Diverse Diseases)
►▼
Show Figures
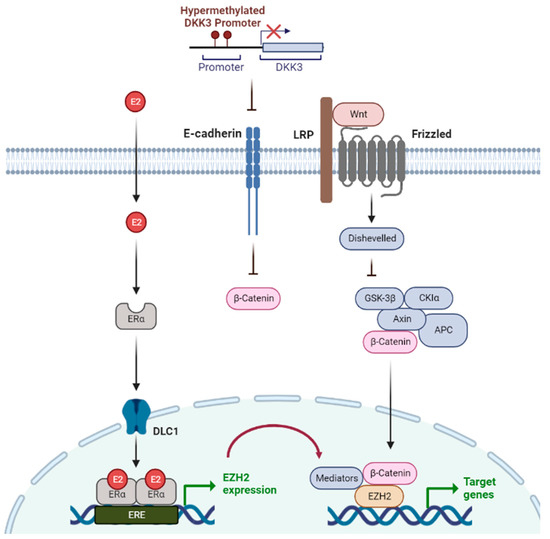
Figure 1
Open AccessArticle
SNCA Gene Methylation in Parkinson’s Disease and Multiple System Atrophy
Epigenomes 2023, 7(1), 5; https://doi.org/10.3390/epigenomes7010005 - 06 Feb 2023
Cited by 1
Abstract
In recent years, epigenetic mechanisms have been implicated in the development of multifactorial diseases including neurodegenerative disorders. In Parkinson’s disease (PD), as a synucleinopathy, most studies focused on DNA methylation of SNCA gene coding alpha-synuclein but obtained results were rather contradictory. In another
[...] Read more.
In recent years, epigenetic mechanisms have been implicated in the development of multifactorial diseases including neurodegenerative disorders. In Parkinson’s disease (PD), as a synucleinopathy, most studies focused on DNA methylation of SNCA gene coding alpha-synuclein but obtained results were rather contradictory. In another neurodegenerative synucleinopathy, multiple system atrophy (MSA), very few studies investigated the epigenetic regulation. This study included patients with PD (n = 82), patients with MSA (n = 24), and a control group (n = 50). In three groups, methylation levels of CpG and non-CpG sites in regulatory regions of the SNCA gene were analyzed. We revealed hypomethylation of CpG sites in the SNCA intron 1 in PD and hypermethylation of predominantly non-CpG sites in the SNCA promoter region in MSA. In PD patients, hypomethylation in the intron 1 was associated with earlier age at the disease onset. In MSA patients, hypermethylation in the promotor was associated with shorter disease duration (before examination). These results showed different patterns of the epigenetic regulation in two synucleinopathies—PD and MSA.
Full article
(This article belongs to the Special Issue Non-CpG Methylation)
►▼
Show Figures
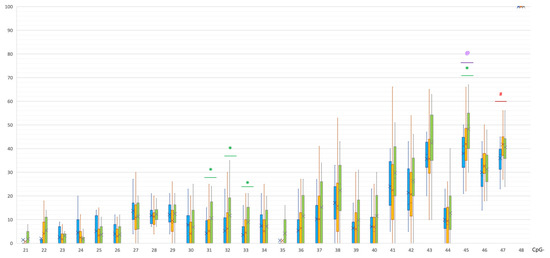
Figure 1
Open AccessArticle
DNA Methylation Is a Potential Biomarker for Cardiometabolic Health in Mexican Children and Adolescents
by
, , , , , , , , and
Epigenomes 2023, 7(1), 4; https://doi.org/10.3390/epigenomes7010004 - 03 Feb 2023
Abstract
DNA methylation (DNAm) is a plausible mechanism underlying cardiometabolic abnormalities, but evidence is limited among youth. This analysis included 410 offspring of the Early Life Exposure in Mexico to Environmental Toxicants (ELEMENT) birth cohort followed up to two time points in late childhood/adolescence.
[...] Read more.
DNA methylation (DNAm) is a plausible mechanism underlying cardiometabolic abnormalities, but evidence is limited among youth. This analysis included 410 offspring of the Early Life Exposure in Mexico to Environmental Toxicants (ELEMENT) birth cohort followed up to two time points in late childhood/adolescence. At Time 1, DNAm was quantified in blood leukocytes at long interspersed nuclear elements (LINE-1), H19, and 11β-hydroxysteroid dehydrogenase type 2 (11β-HSD-2), and at Time 2 in peroxisome proliferator-activated receptor alpha (PPAR-α). At each time point, cardiometabolic risk factors were assessed including lipid profiles, glucose, blood pressure, and anthropometry. Linear mixed effects models were used for LINE-1, H19, and 11β-HSD-2 to account for the repeated-measure outcomes. Linear regression models were conducted for the cross-sectional association between PPAR-α with the outcomes. DNAm at LINE-1 was associated with log glucose at site 1 [β = −0.029, p = 0.0006] and with log high-density lipoprotein cholesterol at site 3 [β = 0.063, p = 0.0072]. 11β-HSD-2 DNAm at site 4 was associated with log glucose (β = −0.018, p = 0.0018). DNAm at LINE-1 and 11β-HSD-2 was associated with few cardiometabolic risk factors among youth in a locus-specific manner. These findings underscore the potential for epigenetic biomarkers to increase our understanding of cardiometabolic risk earlier in life.
Full article
(This article belongs to the Special Issue Environmental Epigenomes)
►▼
Show Figures
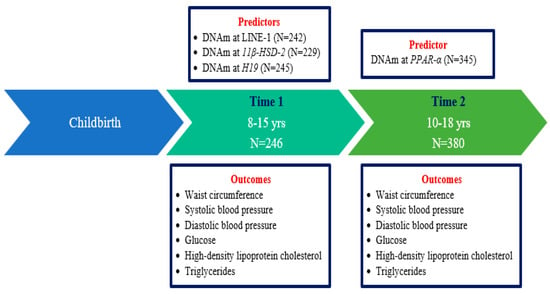
Figure 1
Open AccessEditorial
Acknowledgment to the Reviewers of Epigenomes in 2022
Epigenomes 2023, 7(1), 3; https://doi.org/10.3390/epigenomes7010003 - 13 Jan 2023
Abstract
High-quality academic publishing is built on rigorous peer review [...]
Full article
Open AccessBrief Report
Sex-Specific miRNA Differences in Liquid Biopsies from Subjects with Solid Tumors and Healthy Controls
Epigenomes 2023, 7(1), 2; https://doi.org/10.3390/epigenomes7010002 - 10 Jan 2023
Cited by 3
Abstract
►▼
Show Figures
Dysregulation of epigenetic mechanisms has been recognized to play a crucial role in cancer development, but these mechanisms vary between sexes. Therefore, we focused on sex-specific differences in the context of cancer-based data from a recent study. A total of 12 cell-free DNA
[...] Read more.
Dysregulation of epigenetic mechanisms has been recognized to play a crucial role in cancer development, but these mechanisms vary between sexes. Therefore, we focused on sex-specific differences in the context of cancer-based data from a recent study. A total of 12 cell-free DNA methylation targets in CpG-rich promoter regions and 48 miRNAs were analyzed by qPCR in plasma samples from 8 female and 7 male healthy controls as well as 48 female and 80 male subjects with solid tumors of the bladder, brain, colorectal region (CRC), lung, stomach, pancreas, and liver. Due to the small sample size in some groups and/or the non-balanced distribution of men and women, sex-specific differences were evaluated statistically only in healthy subjects, CRC, stomach or pancreas cancer patients, and all cancer subjects combined (n female/male—8/7, 14/14, 8/15, 6/6, 48/80, respectively). Several miRNAs with opposing expressions between the sexes were observed for healthy subjects (miR-17-5p, miR-26b-5p); CRC patients (miR-186-5p, miR-22-3p, miR-22-5p, miR-25-3p, miR-92a-3p, miR-16-5p); stomach cancer patients (miR-133a-3p, miR-22-5p); and all cancer patients combined (miR-126-3p, miR-21-5p, miR-92a-3p, miR-183-5p). Moreover, sex-specific correlations that were dependent on cancer stage were observed in women (miR-27a-3p) and men (miR-17-5p, miR-20a-5p). Our results indicate the complex and distinct role of epigenetic regulation, particularly miRNAs, depending not only on the health status but also on the sex of the patient. The same miRNAs could have diverse effects in different tissues and opposing effects between the biological sexes, which should be considered in biomarker research.
Full article
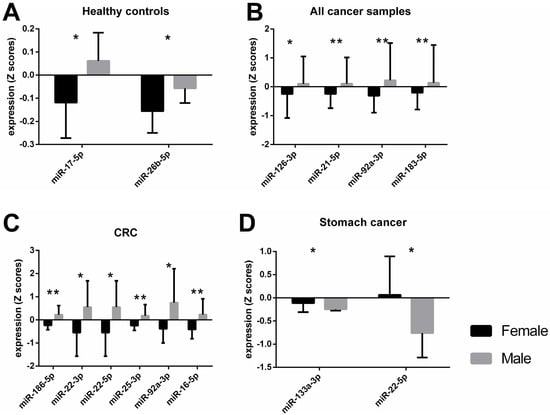
Figure 1
Open AccessReview
Environmental Adaptation of Genetically Uniform Organisms with the Help of Epigenetic Mechanisms—An Insightful Perspective on Ecoepigenetics
by
Epigenomes 2023, 7(1), 1; https://doi.org/10.3390/epigenomes7010001 - 26 Dec 2022
Abstract
Organisms adapt to different environments by selection of the most suitable phenotypes from the standing genetic variation or by phenotypic plasticity, the ability of single genotypes to produce different phenotypes in different environments. Because of near genetic identity, asexually reproducing populations are particularly
[...] Read more.
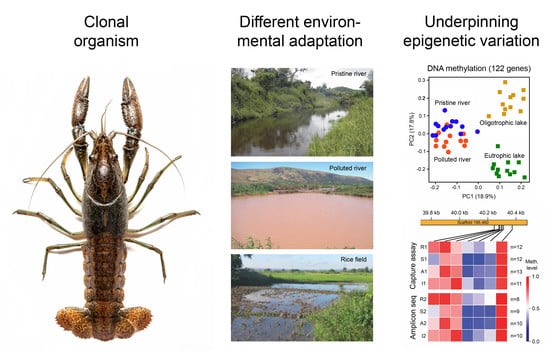
Organisms adapt to different environments by selection of the most suitable phenotypes from the standing genetic variation or by phenotypic plasticity, the ability of single genotypes to produce different phenotypes in different environments. Because of near genetic identity, asexually reproducing populations are particularly suitable for the investigation of the potential and molecular underpinning of the latter alternative in depth. Recent analyses on the whole-genome scale of differently adapted clonal animals and plants demonstrated that epigenetic mechanisms such as DNA methylation, histone modifications and non-coding RNAs are among the molecular pathways supporting phenotypic plasticity and that epigenetic variation is used to stably adapt to different environments. Case studies revealed habitat-specific epigenetic fingerprints that were maintained over subsequent years pointing at the existence of epigenetic ecotypes. Environmentally induced epimutations and corresponding gene expression changes provide an ideal means for fast and directional adaptation to changing or new conditions, because they can synchronously alter phenotypes in many population members. Because microorganisms inclusive of human pathogens also exploit epigenetically mediated phenotypic variation for environmental adaptation, this phenomenon is considered a universal biological principle. The production of different phenotypes from the same DNA sequence in response to environmental cues by epigenetic mechanisms also provides a mechanistic explanation for the “general-purpose genotype hypothesis” and the “genetic paradox of invasions”.
Full article
(This article belongs to the Special Issue Environmental Epigenomes)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage
by
, , , , , and
Epigenomes 2022, 6(4), 43; https://doi.org/10.3390/epigenomes6040043 - 09 Dec 2022
Abstract
TBX15, which encodes a differentiation-related transcription factor, displays promoter-adjacent DNA hypermethylation in myoblasts and skeletal muscle (psoas) that is absent from non-expressing cells in other lineages. By whole-genome bisulfite sequencing (WGBS) and enzymatic methyl-seq (EM-seq), these hypermethylated regions were found to border
[...] Read more.
TBX15, which encodes a differentiation-related transcription factor, displays promoter-adjacent DNA hypermethylation in myoblasts and skeletal muscle (psoas) that is absent from non-expressing cells in other lineages. By whole-genome bisulfite sequencing (WGBS) and enzymatic methyl-seq (EM-seq), these hypermethylated regions were found to border both sides of a constitutively unmethylated promoter. To understand the functionality of this DNA hypermethylation, we cloned the differentially methylated sequences (DMRs) in CpG-free reporter vectors and tested them for promoter or enhancer activity upon transient transfection. These cloned regions exhibited strong promoter activity and, when placed upstream of a weak promoter, strong enhancer activity specifically in myoblast host cells. In vitro CpG methylation targeted to the DMR sequences in the plasmids resulted in 86–100% loss of promoter or enhancer activity, depending on the insert sequence. These results as well as chromatin epigenetic and transcription profiles for this gene in various cell types support the hypothesis that DNA hypermethylation immediately upstream and downstream of the unmethylated promoter region suppresses enhancer/extended promoter activity, thereby downmodulating, but not silencing, expression in myoblasts and certain kinds of skeletal muscle. This promoter-border hypermethylation was not found in cell types with a silent TBX15 gene, and these cells, instead, exhibit repressive chromatin in and around the promoter. TBX18, TBX2, TBX3 and TBX1 display TBX15-like hypermethylated DMRs at their promoter borders and preferential expression in myoblasts. Therefore, promoter-adjacent DNA hypermethylation for downmodulating transcription to prevent overexpression may be used more frequently for transcription regulation than currently appreciated.
Full article
(This article belongs to the Special Issue Recent Advances in Biological Methylation 2022)
►▼
Show Figures
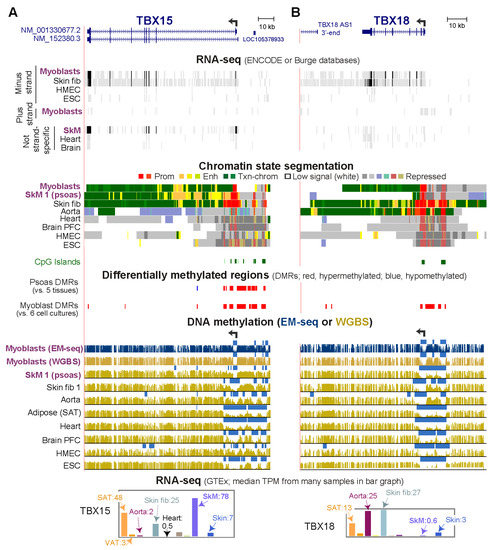
Figure 1
Highly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Cells, Epigenomes, Genes, IJMS, IJTM
Stem Cell Differentiation and Applications
Topic Editors: Hiroyuki Hirai, Haiyun Pei, Atsushi AsakuraDeadline: 20 October 2023
Topic in
Biology, Epigenomes, Societies, Life, Healthcare
Biosocial Studies
Topic Editors: Juan R. Coca, Adolfo Cordero-Rivera, Isabel Castro-PiedrasDeadline: 21 November 2023

Conferences
Special Issues
Special Issue in
Epigenomes
X-Chromosome Inactivation
Guest Editors: Vincent Pasque, Bradley Philip BalatonDeadline: 30 September 2023
Special Issue in
Epigenomes
New Insights into Epigenetics and Metabolism
Guest Editors: Che-Kun James Shen, Guoping FanDeadline: 20 November 2023
Special Issue in
Epigenomes
Environmental Epigenomes
Guest Editors: Frédéric Silvestre, Bambarendage PereraDeadline: 31 December 2023
Topical Collections
Topical Collection in
Epigenomes
Epigenetic Mechanisms in Diabetes Research
Collection Editor: Assam El-Osta
Topical Collection in
Epigenomes
Feature Papers in Epigenomes
Collection Editors: Ivana De la Serna, Che-Kun James Shen


















