-
 Reduced Binding between Omicron B.1.1.529 and the Human ACE2 Receptor in a Surrogate Virus Neutralization Test for SARS-CoV-2
Reduced Binding between Omicron B.1.1.529 and the Human ACE2 Receptor in a Surrogate Virus Neutralization Test for SARS-CoV-2 -
 Comparing Heterologous and Homologous COVID-19 Vaccination: A Longitudinal Study of Antibody Decay
Comparing Heterologous and Homologous COVID-19 Vaccination: A Longitudinal Study of Antibody Decay -
 Microchimerism in Baboon Recipients of Pig Hearts
Microchimerism in Baboon Recipients of Pig Hearts -
 Dengue Virus Infection of Human Retinal Cells
Dengue Virus Infection of Human Retinal Cells -
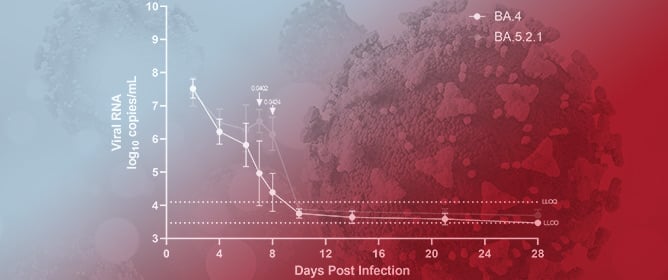 The Omicron Sub-Variant BA.4 Displays a Remarkable Lack of Clinical Signs in a Golden Syrian Hamster Model of SARS-CoV-2 Infection
The Omicron Sub-Variant BA.4 Displays a Remarkable Lack of Clinical Signs in a Golden Syrian Hamster Model of SARS-CoV-2 Infection
Journal Description
Viruses
Viruses
is a peer-reviewed, open access journal of virology, published monthly online by MDPI. The American Society for Virology (ASV), Spanish Society for Virology (SEV), Canadian Society for Virology (CSV), Italian Society for Virology (SIV-ISV), Australasian Virology Society (AVS) and others are affiliated with Viruses and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, MEDLINE, PMC, Embase, PubAg, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Virology) / CiteScore - Q1 (Infectious Diseases)
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.8 days after submission; acceptance to publication is undertaken in 2.5 days (median values for papers published in this journal in the first half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Companion journals for Viruses include: COVID and Zoonotic Diseases.
Impact Factor:
4.7 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
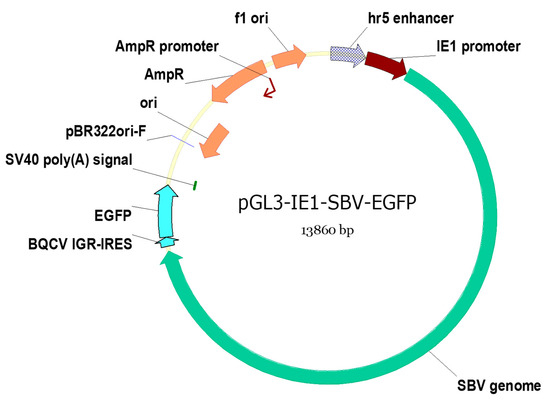
A DNA Plasmid-Based Approach for Efficient Synthesis of Sacbrood Virus Infectious Clones within Host Cells
Viruses 2023, 15(9), 1866; https://doi.org/10.3390/v15091866 (registering DOI) - 01 Sep 2023
Abstract
RNA viruses are often cited as a significant factor affecting the populations of both domestic honey bees and wild pollinators. To expedite the development of effective countermeasures against these viruses, a more comprehensive understanding of virus biology necessitates extensive collaboration among scientists from
[...] Read more.
RNA viruses are often cited as a significant factor affecting the populations of both domestic honey bees and wild pollinators. To expedite the development of effective countermeasures against these viruses, a more comprehensive understanding of virus biology necessitates extensive collaboration among scientists from diverse research fields. While the infectious virus clone is a robust tool for studying virus diseases, the current methods for synthesizing infectious clones of bee-infecting RNA viruses entail the in vitro transcription of the viral genome RNA in 8–10 kb, presenting challenges in reproducibility and distribution. This article reports on the synthesis of an infectious clone of the Chinese variant sacbrood virus (SBV) using a DNA plasmid containing an Autographa californica multiple nucleopolyhedrovirus (AcMNPV) immediate-early protein (IE1) promoter to trigger transcription of the downstream viral genome within hosts. The results demonstrate that the IE1-SBV plasmid can synthesize SBV clones in a widely used lepidopteran immortal cell line (Sf9) and honey bee pupae. Furthermore, the negative strand of the clone was detected in both Sf9 cells and honey bee pupae, indicating active infection and replication. However, the transfection of Sf9 cells was observed in only a limited proportion (less than 10%) of the cells, and the infection did not appear to spread to adjacent cells or form infective virions. The injection of honey bee pupae with 2500 ng of the IE1-SBV plasmid resulted in high infection rates in Apis cerana pupae but low rates in A. mellifera pupae, although the dosage was comparatively high compared with other studies using in vitro transcribed viral RNA. Our findings suggest that the synthesis of bee-infecting RNA viruses using DNA plasmids is feasible, albeit requiring additional optimization. However, this method holds substantial potential for facilitating the production of clones with various sequence modifications, enabling the exploration of viral gene functions and biology. The ease of distributing infectious clones in DNA plasmid form may foster collaboration among scientists in applying the clone to bee biology, ecology, and behavior, ultimately offering a comprehensive approach to managing virus diseases in the future.
Full article
(This article belongs to the Section Insect Viruses)
►
Show Figures
Open AccessBrief Report
Detection of H5N1 High Pathogenicity Avian Influenza Viruses in Four Raptors and Two Geese in Japan in the Fall of 2022
by
, , , , , , , , and
Viruses 2023, 15(9), 1865; https://doi.org/10.3390/v15091865 (registering DOI) - 01 Sep 2023
Abstract
In the fall of 2022, high pathogenicity avian influenza viruses (HPAIVs) were detected from raptors and geese in Japan, a month earlier than in past years, indicating a shift in detection patterns. In this study, we conducted a phylogenetic analysis on H5N1 HPAIVs
[...] Read more.
In the fall of 2022, high pathogenicity avian influenza viruses (HPAIVs) were detected from raptors and geese in Japan, a month earlier than in past years, indicating a shift in detection patterns. In this study, we conducted a phylogenetic analysis on H5N1 HPAIVs detected from six wild birds during the 2022/2023 season to determine their genetic origins. Our findings revealed that these HPAIVs belong to the G2 group within clade 2.3.4.4b, with all isolates classified into three subgroups: G2b, G2d, and G2c. The genetic background of the G2b virus (a peregrine falcon-derived strain) and G2d viruses (two raptors and two geese-derived strains) were the same as those detected in Japan in the 2021/2022 season. Since no HPAI cases were reported in Japan during the summer of 2022, it is probable that migratory birds reintroduced the G2b and G2d viruses. Conversely, the G2c virus (a raptor-derived strain) was first recognized in Japan in the fall of 2022. This strain might share a common ancestor with HPAIVs from Asia and West Siberia observed in the 2021/2022 season. The early migration of waterfowl to Japan in the fall of 2022 could have facilitated the early invasion of HPAIVs.
Full article
(This article belongs to the Special Issue Avian Respiratory Viruses, Volume III)
►▼
Show Figures
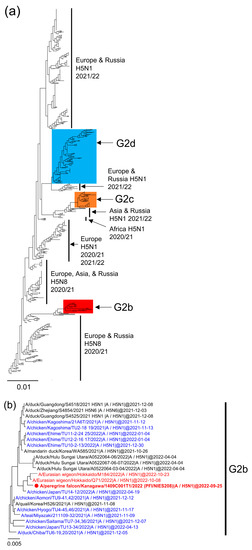
Figure 1
Open AccessArticle
Human Rotavirus Replicates in Salivary Glands and Primes Immune Responses in Facial and Intestinal Lymphoid Tissues of Gnotobiotic Pigs
by
, , , , , , , , , , , , and
Viruses 2023, 15(9), 1864; https://doi.org/10.3390/v15091864 (registering DOI) - 31 Aug 2023
Abstract
Human rotavirus (HRV) is a leading cause of viral gastroenteritis in children across the globe. The virus has long been established as a pathogen of the gastrointestinal tract, targeting small intestine epithelial cells and leading to diarrhea, nausea, and vomiting. Recently, this classical
[...] Read more.
Human rotavirus (HRV) is a leading cause of viral gastroenteritis in children across the globe. The virus has long been established as a pathogen of the gastrointestinal tract, targeting small intestine epithelial cells and leading to diarrhea, nausea, and vomiting. Recently, this classical infection pathway was challenged by the findings that murine strains of rotavirus can infect the salivary glands of pups and dams and transmit via saliva from pups to dams during suckling. Here, we aimed to determine if HRV was also capable of infecting salivary glands and spreading in saliva using a gnotobiotic (Gn) pig model of HRV infection and disease. Gn pigs were orally inoculated with various strains of HRV, and virus shedding was monitored for several days post-inoculation. HRV was shed nasally and in feces in all inoculated pigs. Infectious HRV was detected in the saliva of four piglets. Structural and non-structural HRV proteins, as well as the HRV genome, were detected in the intestinal and facial tissues of inoculated pigs. The pigs developed high IgM antibody responses in serum and small intestinal contents at 10 days post-inoculation. Additionally, inoculated pigs had HRV-specific IgM antibody-secreting cells present in the ileum, tonsils, and facial lymphoid tissues. Taken together, these findings indicate that HRV can replicate in salivary tissues and prime immune responses in both intestinal and facial lymphoid tissues of Gn pigs.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures
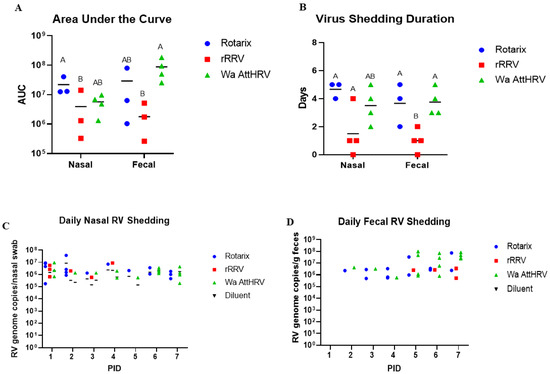
Figure 1
Open AccessArticle
A CRISPR Screen of HIV Dependency Factors Reveals That CCNT1 Is Non-Essential in T Cells but Required for HIV-1 Reactivation from Latency
Viruses 2023, 15(9), 1863; https://doi.org/10.3390/v15091863 (registering DOI) - 31 Aug 2023
Abstract
We sought to explore the hypothesis that host factors required for HIV-1 replication also play a role in latency reversal. Using a CRISPR gene library of putative HIV dependency factors, we performed a screen to identify genes required for latency reactivation. We identified
[...] Read more.
We sought to explore the hypothesis that host factors required for HIV-1 replication also play a role in latency reversal. Using a CRISPR gene library of putative HIV dependency factors, we performed a screen to identify genes required for latency reactivation. We identified several HIV-1 dependency factors that play a key role in HIV-1 latency reactivation including ELL, UBE2M, TBL1XR1, HDAC3, AMBRA1, and ALYREF. The knockout of Cyclin T1 (CCNT1), a component of the P-TEFb complex that is important for transcription elongation, was the top hit in the screen and had the largest effect on HIV latency reversal with a wide variety of latency reversal agents. Moreover, CCNT1 knockout prevents latency reactivation in a primary CD4+ T cell model of HIV latency without affecting the activation of these cells. RNA sequencing data showed that CCNT1 regulates HIV-1 proviral genes to a larger extent than any other host gene and had no significant effects on RNA transcripts in primary T cells after activation. We conclude that CCNT1 function is non-essential in T cells but is absolutely required for HIV latency reversal.
Full article
(This article belongs to the Special Issue Regulation of HIV-1 Transcription and Latency)
►▼
Show Figures
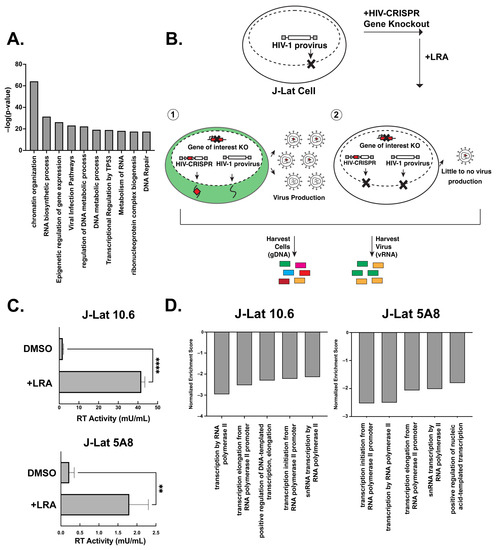
Figure 1
Open AccessBrief Report
The Rise and Fall of Omicron BA.1 Variant as Seen in Wastewater Supports Epidemiological Model Predictions
by
, , , , , , and
Viruses 2023, 15(9), 1862; https://doi.org/10.3390/v15091862 (registering DOI) - 31 Aug 2023
Abstract
The COVID-19 pandemic caused by the SARS-CoV-2 virus has inflicted significant mortality and morbidity worldwide. Continuous virus mutations have led to the emergence of new variants. The Omicron BA.1 sub-lineage prevailed as the dominant variant globally at the beginning of 2022 but was
[...] Read more.
The COVID-19 pandemic caused by the SARS-CoV-2 virus has inflicted significant mortality and morbidity worldwide. Continuous virus mutations have led to the emergence of new variants. The Omicron BA.1 sub-lineage prevailed as the dominant variant globally at the beginning of 2022 but was subsequently replaced by BA.2 in numerous countries. Wastewater-based epidemiology (WBE) offers an efficient tool for capturing viral shedding from infected individuals, enabling early detection of potential pandemic outbreaks without relying solely on community cooperation and clinical testing resources. This study integrated RT-qPCR assays for detecting general SARS-CoV-2 and its variants levels in wastewater into a modified triple susceptible-infected-recovered-susceptible (SIRS) model. The emergence of the Omicron BA.1 variant was observed, replacing the presence of its predecessor, the Delta variant. Comparative analysis between the wastewater data and the modified SIRS model effectively described the BA.1 and subsequent BA.2 waves, with the decline of the Delta variant aligning with its diminished presence below the detection threshold in wastewater. This study demonstrates the potential of WBE as a valuable tool for future pandemics. Furthermore, by analyzing the sensitivity of different variants to model parameters, we are able to deduce real-life values of cross-variant immunity probabilities, emphasizing the asymmetry in their strength.
Full article
(This article belongs to the Special Issue Wastewater-Based Epidemiology (WBE) in COVID-19 Pandemics)
►▼
Show Figures
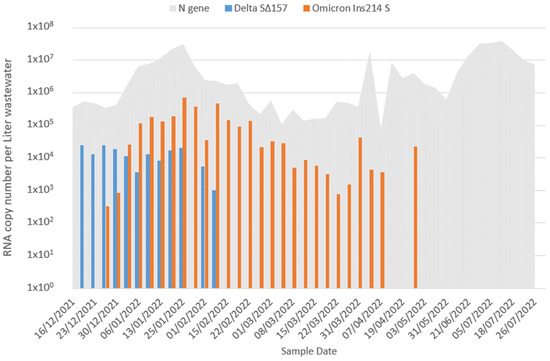
Figure 1
Open AccessReview
Global Burden of Lumpy Skin Disease, Outbreaks, and Future Challenges
by
, , , , , and
Viruses 2023, 15(9), 1861; https://doi.org/10.3390/v15091861 (registering DOI) - 31 Aug 2023
Abstract
Lumpy skin disease (LSD), a current global concern, causes economic devastation in livestock industries, with cattle and water buffalo reported to have higher morbidity and lower mortality rates. LSD is caused by lumpy skin disease virus (LSDV), a member of the Poxviridae family.
[...] Read more.
Lumpy skin disease (LSD), a current global concern, causes economic devastation in livestock industries, with cattle and water buffalo reported to have higher morbidity and lower mortality rates. LSD is caused by lumpy skin disease virus (LSDV), a member of the Poxviridae family. It is an enzootic, rapidly explorative and sometimes fatal infection, characterized by multiple raised nodules on the skin of infected animals. It was first reported in Zambia in 1929 and is considered endemic in Africa south of the Sahara desert. It has gradually spread beyond Africa into the Middle East, with periodic occurrences in Asian and East European countries. Recently, it has been spreading in most Asian countries including far East Asia and threatens incursion to LSD-free countries. Rapid and accurate diagnostic capabilities, virus identification, vaccine development, vector control, regional and international collaborations and effective biosecurity policies are important for the control, prevention, and eradication of LSD infections. This review critically evaluates the global burden of LSD, the chronological historical outbreaks of LSD, and future directions for collaborative global actions.
Full article
(This article belongs to the Special Issue Capripox Viruses: A Continuing Transboundary Threat to Animal Health)
►▼
Show Figures
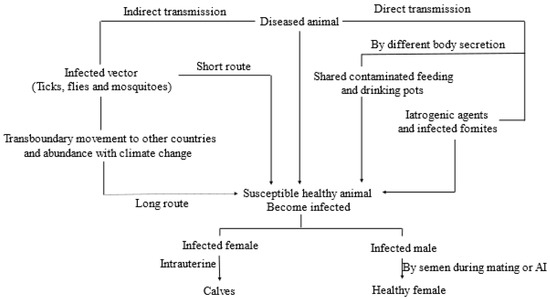
Figure 1
Open AccessReview
Viral Co-Infection in Bats: A Systematic Review
Viruses 2023, 15(9), 1860; https://doi.org/10.3390/v15091860 (registering DOI) - 31 Aug 2023
Abstract
Co-infection is an underappreciated phenomenon in contemporary disease ecology despite its ubiquity and importance in nature. Viruses, and other co-infecting agents, can interact in ways that shape host and agent communities, influence infection dynamics, and drive evolutionary selective pressures. Bats are host to
[...] Read more.
Co-infection is an underappreciated phenomenon in contemporary disease ecology despite its ubiquity and importance in nature. Viruses, and other co-infecting agents, can interact in ways that shape host and agent communities, influence infection dynamics, and drive evolutionary selective pressures. Bats are host to many viruses of zoonotic potential and have drawn increasing attention in their role as wildlife reservoirs for human spillover. However, the role of co-infection in driving viral transmission dynamics within bats is unknown. Here, we systematically review peer-reviewed literature reporting viral co-infections in bats. We show that viral co-infection is common in bats but is often only reported as an incidental finding. Biases identified in our study database related to virus and host species were pre-existing in virus studies of bats generally. Studies largely speculated on the role co-infection plays in viral recombination and few investigated potential drivers or impacts of co-infection. Our results demonstrate that current knowledge of co-infection in bats is an ad hoc by-product of viral discovery efforts, and that future targeted co-infection studies will improve our understanding of the role it plays. Adding to the broader context of co-infection studies in other wildlife species, we anticipate our review will inform future co-infection study design and reporting in bats. Consideration of detection strategy, including potential viral targets, and appropriate analysis methodology will provide more robust results and facilitate further investigation of the role of viral co-infection in bat reservoirs.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures

Graphical abstract
Open AccessArticle
Lichen or Associated Micro-Organism Compounds Are Active against Human Coronaviruses
by
, , , , , , , , , and
Viruses 2023, 15(9), 1859; https://doi.org/10.3390/v15091859 - 31 Aug 2023
Abstract
(1) Background: Since the emergence of SARS-CoV-2, responsible for the COVID-19 pandemic, efforts have been made to identify antiviral compounds against human coronaviruses. With the aim of increasing the diversity of molecule scaffolds, 42 natural compounds, of which 28 were isolated from lichens
[...] Read more.
(1) Background: Since the emergence of SARS-CoV-2, responsible for the COVID-19 pandemic, efforts have been made to identify antiviral compounds against human coronaviruses. With the aim of increasing the diversity of molecule scaffolds, 42 natural compounds, of which 28 were isolated from lichens and 14 from their associated microorganisms (bacteria and fungi), were screened against human coronavirus HCoV-229E. (2) Methods: Antiviral assays were performed using HCoV-229E in Huh-7 and Huh-7/TMPRSS2 cells and SARS-CoV-2 in a Vero-81-derived clone with a GFP reporter probe. (3) Results: Four lichen compounds, including chloroatranol, emodin, perlatolic acid and vulpinic acid, displayed high activities against HCoV-229E (IC50 = 68.86, 59.25, 16.42 and 14.58 μM, respectively) and no toxicity at active concentrations. Kinetics studies were performed to determine their mode of action. The four compounds were active when added at the replication step. Due to their significant activity, they were further tested on SARS-CoV-2. Perlatolic acid was shown to be active against SARS-CoV-2. (4) Conclusions: Taken together, these results show that lichens are a source of interesting antiviral agents against human coronaviruses. Moreover, perlatolic acid might be further studied for its pan-coronavirus antiviral activity.
Full article
(This article belongs to the Special Issue Novel Antiviral Agents: Synthesis, Molecular Modelling Studies and Biological Investigation 2nd Edition)
►▼
Show Figures
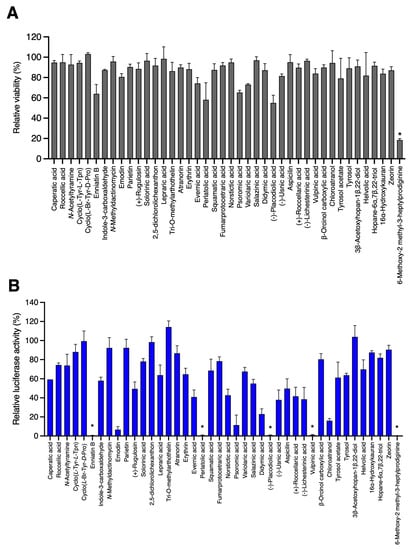
Figure 1
Open AccessArticle
Carambolaside W Inhibited H1N1 Influenza Virus-Induced Oxidative Stress through STAT-3/BCL-XL Signaling Pathway
by
, , , , , , , and
Viruses 2023, 15(9), 1858; https://doi.org/10.3390/v15091858 - 31 Aug 2023
Abstract
The H1N1 influenza virus is highly infectious and pathogenic, and in recent years, it has often presented seasonal mass outbreaks of infection. People infected with H1N1 will develop a high fever and other respiratory infection symptoms. If not treated in time, complications such
[...] Read more.
The H1N1 influenza virus is highly infectious and pathogenic, and in recent years, it has often presented seasonal mass outbreaks of infection. People infected with H1N1 will develop a high fever and other respiratory infection symptoms. If not treated in time, complications such as pneumonia may occur. In this study, we focused on developing drugs that can effectively fight against with H1N1 virus. A flavonoid glycoside was extracted from the carambola, then characterized by HR-ESI-MS with the molecular formula C47H58O2, and named carambolaside W. The flavonoid glycosides were found to have good anti-H1N1 influenza virus effects. In this study, we verified that carambolaside W has low toxicity and can effectively inhibit influenza virus replication in vitro. H1N1 virus infection induces intracellular oxidative stress damage to accelerate disease progression. The results showed that carambolaside W effectively inhibited the oxidative stress caused by H1N1 infection. The Western blot assay also revealed that carambolaside W alters the expression of apoptosis-related proteins in vitro and exerts a good anti-H1N1 influenza virus effect. In summary, carambolaside W is a low-toxicity natural flavonoid that can effectively treat the H1N1 influenza virus as a potential anti-H1N1 virus agent.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures
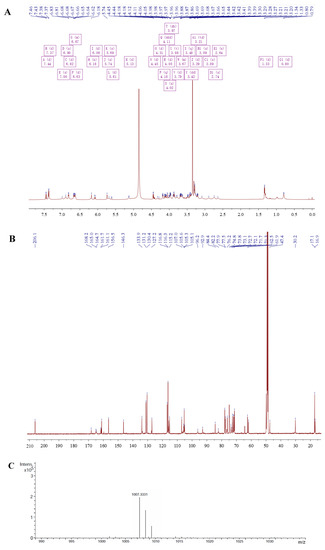
Figure 1
Open AccessCommunication
Burden of HPV-Related Hospitalization in Germany from 2000 to 2021
Viruses 2023, 15(9), 1857; https://doi.org/10.3390/v15091857 - 31 Aug 2023
Abstract
HPV has been linked to the development of precancerous and cancerous lesions. The aim of this study was to evaluate the burden of HPV-related hospitalization in Germany from 2000 to 2021 and the potential impact of the COVID-19 pandemic on it. Methods: We
[...] Read more.
HPV has been linked to the development of precancerous and cancerous lesions. The aim of this study was to evaluate the burden of HPV-related hospitalization in Germany from 2000 to 2021 and the potential impact of the COVID-19 pandemic on it. Methods: We performed a retrospective query using data from the German Statistical Office from 2000 to 2021, including hospital admission, inpatient mortality and hospital stay length data on cervical cancer/dysplasia, female genitourinary tract, anal, penile, head and neck cancers. Results: The HPV-attributable hospitalization rate per 100,000 inhabitants in Germany has decreased over time, from 89 cases in 2000 to 60 in 2021, with an average annual percent change (AAPC) of −1.93 (CI −2.08–−1.79, p < 0.05). The same trend was observed for the average hospital stay, which declined from 9 to 7 days, with an AAPC of −1.33 (CI −1.52–−1.21, p < 0.05). An undulating but overall slightly declining pattern was observed for the inpatient mortality (AAPC −0.92, CI −1.21–−0.64, p < 0.05). We observed a reduction in the hospitalization rates for invasive and non-invasive cervical cancer, which was observed in almost all age groups and in all German federal states. Conclusion: Our study provides a comprehensive analysis of the trends in HPV-related hospitalizations over the past two decades. The decline in hospitalization rates for cervical cancer and dysplasia suggests the potential efficacy of the HPV vaccination and screening programs.
Full article
(This article belongs to the Special Issue Opportunistic Viral Infections)
►▼
Show Figures
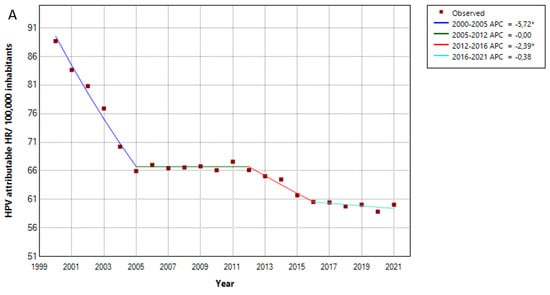
Figure 1
Open AccessReview
Envelope Recombination: A Major Driver in Shaping Retroviral Diversification and Evolution within the Host Genome
Viruses 2023, 15(9), 1856; https://doi.org/10.3390/v15091856 - 31 Aug 2023
Abstract
Endogenous retroviruses (ERVs) are integrated into host DNA as the result of ancient germ line infections, primarily by extinct exogenous retroviruses. Thus, vertebrates’ genomes contain thousands of ERV copies, providing a “fossil” record for ancestral retroviral diversity and its evolution within the host
[...] Read more.
Endogenous retroviruses (ERVs) are integrated into host DNA as the result of ancient germ line infections, primarily by extinct exogenous retroviruses. Thus, vertebrates’ genomes contain thousands of ERV copies, providing a “fossil” record for ancestral retroviral diversity and its evolution within the host genome. Like other retroviruses, the ERV proviral sequence consists of gag, pro, pol, and env genes flanked by long terminal repeats (LTRs). Particularly, the env gene encodes for the envelope proteins that initiate the infection process by binding to the host cellular receptor(s), causing membrane fusion. For this reason, a major element in understanding ERVs’ evolutionary trajectory is the characterization of env changes over time. Most of the studies dedicated to ERVs’ env have been aimed at finding an “actual” physiological or pathological function, while few of them have focused on how these genes were once acquired and modified within the host. Once acquired into the organism, genome ERVs undergo common cellular events, including recombination. Indeed, genome recombination plays a role in ERV evolutionary dynamics. Retroviral recombination events that might have been involved in env divergence include the acquisition of env genes from distantly related retroviruses, env swapping facilitating multiple cross-species transmission over millions of years, ectopic recombination between the homologous sequences present in different positions in the chromosomes, and template switching during transcriptional events. The occurrence of these recombinational events might have aided in shaping retroviral diversification and evolution until the present day. Hence, this review describes and discusses in detail the reported recombination events involving ERV env to provide the basis for further studies in the field.
Full article
(This article belongs to the Special Issue Endogenous Retrovirus Proteins and Their Functions)
►▼
Show Figures
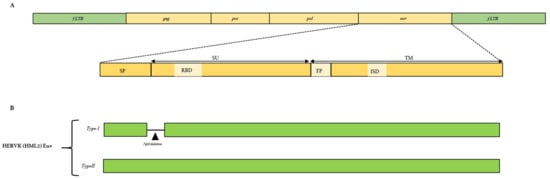
Figure 1
Open AccessArticle
An Integrated Research–Clinical BSL-2 Platform for a Live SARS-CoV-2 Neutralization Assay
by
, , , , , , , , , , , , , and
Viruses 2023, 15(9), 1855; https://doi.org/10.3390/v15091855 - 31 Aug 2023
Abstract
A reliable and efficient serological test is crucial for monitoring neutralizing antibodies against SARS-CoV-2 and its variants of concern (VOCs). Here, we present an integrated research–clinical platform for a live SARS-CoV-2 neutralization assay, utilizing highly attenuated SARS-CoV-2 (Δ3678_WA1-spike). This strain contains mutations in
[...] Read more.
A reliable and efficient serological test is crucial for monitoring neutralizing antibodies against SARS-CoV-2 and its variants of concern (VOCs). Here, we present an integrated research–clinical platform for a live SARS-CoV-2 neutralization assay, utilizing highly attenuated SARS-CoV-2 (Δ3678_WA1-spike). This strain contains mutations in viral transcription regulation sequences and deletion in the open-reading-frames 3, 6, 7, and 8, allowing for safe handling in biosafety level 2 (BSL-2) laboratories. Building on this backbone, we constructed a genetically stable reporter virus (mGFP Δ3678_WA1-spike) by incorporating a modified green fluorescent protein sequence (mGFP). We also constructed mGFP Δ3678_BA.5-spike and mGFP Δ3678_XBB.1.5-spike by substituting the WA1 spike with variants BA.5 and XBB.1.5 spike, respectively. All three viruses exhibit robust fluorescent signals in infected cells and neutralization titers in an optimized fluorescence reduction neutralization assay that highly correlates with a conventional plaque reduction assay. Furthermore, we established that a streamlined robot-aided Bench-to-Clinics COVID-19 Neutralization Test workflow demonstrated remarkably sensitive, specific, reproducible, and accurate characteristics, allowing the assessment of neutralization titers against SARS-CoV-2 variants within 24 h after sample receiving. Overall, our innovative approach provides a valuable avenue for large-scale testing of clinical samples against SARS-CoV-2 and VOCs at BSL-2, supporting pandemic preparedness and response strategies.
Full article
(This article belongs to the Section SARS-CoV-2 and COVID-19)
►▼
Show Figures
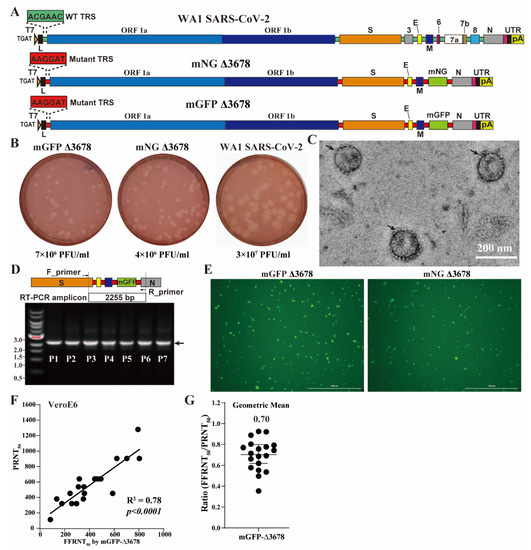
Figure 1
Open AccessPerspective
Host Membranes as Drivers of Virus Evolution
Viruses 2023, 15(9), 1854; https://doi.org/10.3390/v15091854 - 31 Aug 2023
Abstract
The molecular mechanisms controlling the adaptation of viruses to host cells are generally poorly documented. An essential issue to resolve is whether host membranes, and especially lipid rafts, which are usually considered passive gateways for many enveloped viruses, also encode informational guidelines that
[...] Read more.
The molecular mechanisms controlling the adaptation of viruses to host cells are generally poorly documented. An essential issue to resolve is whether host membranes, and especially lipid rafts, which are usually considered passive gateways for many enveloped viruses, also encode informational guidelines that could determine virus evolution. Due to their enrichment in gangliosides which confer an electronegative surface potential, lipid rafts impose a first control level favoring the selection of viruses with enhanced cationic areas, as illustrated by SARS-CoV-2 variants. Ganglioside clusters attract viral particles in a dynamic electrostatic funnel, the more cationic viruses of a viral population winning the race. However, electrostatic forces account for only a small part of the energy of raft-virus interaction, which depends mainly on the ability of viruses to form a network of hydrogen bonds with raft gangliosides. This fine tuning of virus-ganglioside interactions, which is essential to stabilize the virus on the host membrane, generates a second level of selection pressure driven by a typical induced-fit mechanism. Gangliosides play an active role in this process, wrapping around the virus spikes through a dynamic quicksand-like mechanism. Viruses are thus in an endless race for access to lipid rafts, and they are bound to evolve perpetually, combining speed (electrostatic potential) and precision (fine tuning of amino acids) under the selective pressure of the immune system. Deciphering the host membrane guidelines controlling virus evolution mechanisms may open new avenues for the design of innovative antivirals.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
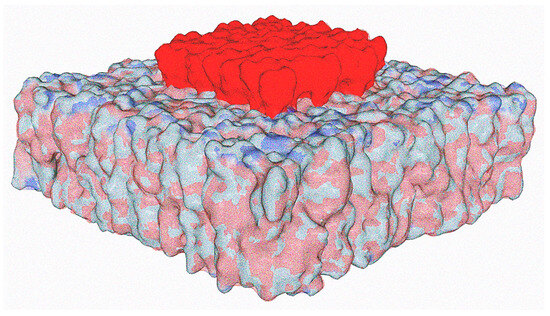
Figure 1
Open AccessArticle
BET Inhibitor JQ1 Attenuates Feline Leukemia Virus DNA, Provirus, and Antigen Production in Domestic Cat Cell Lines
Viruses 2023, 15(9), 1853; https://doi.org/10.3390/v15091853 - 31 Aug 2023
Abstract
Feline leukemia virus (FeLV) is a cosmopolitan gammaretrovirus that causes lifelong infections and fatal diseases, including leukemias, lymphomas, immunodeficiencies, and anemias, in domestic and wild felids. There is currently no definitive treatment for FeLV, and while existing vaccines reduce the prevalence of progressive
[...] Read more.
Feline leukemia virus (FeLV) is a cosmopolitan gammaretrovirus that causes lifelong infections and fatal diseases, including leukemias, lymphomas, immunodeficiencies, and anemias, in domestic and wild felids. There is currently no definitive treatment for FeLV, and while existing vaccines reduce the prevalence of progressive infections, they neither provide sterilizing immunity nor prevent regressive infections that result in viral reservoirs with the potential for reactivation, transmission, and the development of associated clinical diseases. Previous studies of murine leukemia virus (MuLV) established that host cell epigenetic reader bromodomain and extra-terminal domain (BET) proteins facilitate MuLV replication by promoting proviral integration. Here, we provide evidence that this facilitatory effect of BET proteins extends to FeLV. Treatment with the archetypal BET protein bromodomain inhibitor (+)-JQ1 and FeLV challenge of two phenotypically disparate feline cell lines, 81C fibroblasts and 3201 lymphoma cells, significantly reduced FeLV proviral load, total FeLV DNA load, and p27 capsid protein expression at nonlethal concentrations. Moreover, significant decreases in FeLV proviral integration were documented in 81C and 3201 cells. These findings elucidate the importance of BET proteins for efficient FeLV replication, including proviral integration, and provide a potential target for treating FeLV infections.
Full article
(This article belongs to the Section Animal Viruses)
►▼
Show Figures
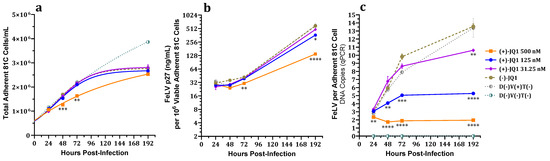
Figure 1
Open AccessArticle
Comprehensive Characterization of Viral Diversity of Female Mosquitoes in Madagascar
by
, , , , , , , , , , , and
Viruses 2023, 15(9), 1852; https://doi.org/10.3390/v15091852 - 31 Aug 2023
Abstract
The diversity and circulation of arboviruses are not much studied in Madagascar. The fact is that arboviral emergences are rarely detected. The existing surveillance system primarily relies on serological detection and records only a few human infections annually. The city of Mahajanga, however,
[...] Read more.
The diversity and circulation of arboviruses are not much studied in Madagascar. The fact is that arboviral emergences are rarely detected. The existing surveillance system primarily relies on serological detection and records only a few human infections annually. The city of Mahajanga, however, experienced a confirmed dengue fever epidemic in 2020 and 2021. This study aimed to characterize and analyze the virome of mosquitoes collected in Mahajanga, near patients with dengue-like syndromes to detect known and unknown viruses as well as investigate the factors contributing to the relative low circulation of arboviruses in the area. A total of 4280 mosquitoes representing at least 12 species from the Aedes, Anopheles, and Culex genera were collected during the dry and the rainy seasons from three sites, following an urbanization gradient. The virome analysis of 2192 female mosquitoes identified a diverse range of viral families and genera and revealed different patterns that are signatures of the influence of the mosquito genus or the season of collection on the composition and abundance of the virome. Despite the absence of known human or veterinary arboviruses, the identification and characterization of viral families, genera, and species in the mosquito virome contribute to our understanding of viral ecology and diversity within mosquito populations in Madagascar. This study serves as a foundation for ongoing surveillance efforts and provides a basis for the development of preventive strategies against various mosquito-borne viral diseases, including known arboviruses.
Full article
(This article belongs to the Section Insect Viruses)
►▼
Show Figures
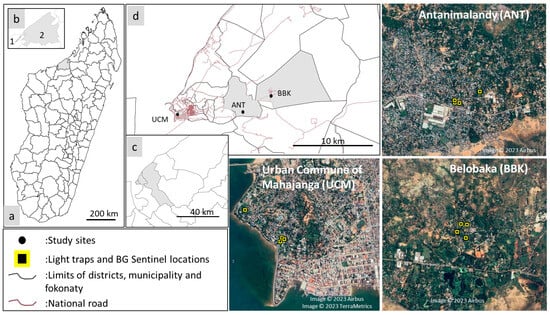
Figure 1
Open AccessArticle
Sequencing and Partial Molecular Characterization of BAB-TMP, the Babeș Strain of the Fixed Rabies Virus Adapted for Multiplication in Cell Lines
by
, , , , , and
Viruses 2023, 15(9), 1851; https://doi.org/10.3390/v15091851 - 31 Aug 2023
Abstract
The rabies virus is a major zoonosis that causes severe nervous disease in humans, leading to paralysis and death. The world’s second anti-rabies center was established in 1888 by Victor Babeș, in Bucharest, where an eponymous strain of rabies was isolated and used
[...] Read more.
The rabies virus is a major zoonosis that causes severe nervous disease in humans, leading to paralysis and death. The world’s second anti-rabies center was established in 1888 by Victor Babeș, in Bucharest, where an eponymous strain of rabies was isolated and used to develop a method for immunization. The Babeș strain of the rabies virus was used for over 100 years in Romania to produce a rabies vaccine for human use, based on animal nerve tissue, thus having a proven history of prophylactic use. The present study aimed to sequence the whole genome of the Babeș strain and to explore its genetic relationships with other vaccine strains as well as to characterize its relevant molecular traits. After being adapted for multiplication in cell lines and designated BAB-TMP, 99% of the viral genome was sequenced. The overall organization of the genome is similar to that of other rabies vaccine strains. Phylogenetic analysis indicated that the BAB-TMP strain is closely related to the Russian RV-97 vaccine strain, and both seem to have a common ancestor. The nucleoprotein gene of the investigated genome was the most conserved, and the glycoprotein showed several unique amino acid substitutions within the major antigenic sites and linear epitopes.
Full article
(This article belongs to the Special Issue Zoonotic Viral Diseases: Drivers, Causes, Prevention and Cure)
►▼
Show Figures
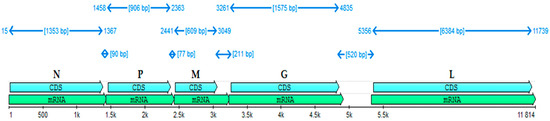
Figure 1
Open AccessArticle
Mining Public Data to Investigate the Virome of Neglected Pollinators and Other Floral Visitors
by
, , , , , , , , , , and
Viruses 2023, 15(9), 1850; https://doi.org/10.3390/v15091850 - 31 Aug 2023
Abstract
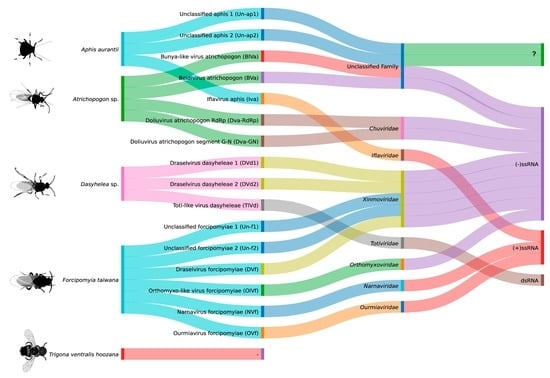
This study reports the virome investigation of pollinator species and other floral visitors associated with plants from the south of Bahia: Aphis aurantii, Atrichopogon sp., Dasyhelea sp., Forcipomyia taiwana, and Trigona ventralis hoozana. Studying viruses in insects associated with economically important crops
[...] Read more.
This study reports the virome investigation of pollinator species and other floral visitors associated with plants from the south of Bahia: Aphis aurantii, Atrichopogon sp., Dasyhelea sp., Forcipomyia taiwana, and Trigona ventralis hoozana. Studying viruses in insects associated with economically important crops is vital to understand transmission dynamics and manage viral diseases that pose as threats for global food security. Using literature mining and public RNA next-generation sequencing data deposited in the NCBI SRA database, we identified potential vectors associated with Malvaceae plant species and characterized the microbial communities resident in these insects. Bacteria and Eukarya dominated the metagenomic analyses of all taxon groups. We also found sequences showing similarity to elements from several viral families, including Bunyavirales, Chuviridae, Iflaviridae, Narnaviridae, Orthomyxoviridae, Rhabdoviridae, Totiviridae, and Xinmoviridae. Phylogenetic analyses indicated the existence of at least 16 new viruses distributed among A. aurantii (3), Atrichopogon sp. (4), Dasyhelea sp. (3), and F. taiwana (6). No novel viruses were found for T. ventralis hoozana. For F. taiwana, the available libraries also allowed us to suggest possible vertical transmission, while for A. aurantii we followed the infection profile along the insect development. Our results highlight the importance of studying the virome of insect species associated with crop pollination, as they may play a crucial role in the transmission of viruses to economically important plants, such as those of the genus Theobroma, or they will reduce the pollination process. This information may be valuable in developing strategies to mitigate the spread of viruses and protect the global industry.
Full article
(This article belongs to the Section Insect Viruses)
►▼
Show Figures
Graphical abstract
Open AccessArticle
Viruses in Laboratory Drosophila and Their Impact on Host Gene Expression
by
and
Viruses 2023, 15(9), 1849; https://doi.org/10.3390/v15091849 - 31 Aug 2023
Abstract
Drosophila melanogaster has one of the best characterized antiviral immune responses among invertebrates. However, relatively few easily transmitted natural virus isolates are available, and so many Drosophila experiments have been performed using artificial infection routes and artificial host–virus combinations. These may not reflect
[...] Read more.
Drosophila melanogaster has one of the best characterized antiviral immune responses among invertebrates. However, relatively few easily transmitted natural virus isolates are available, and so many Drosophila experiments have been performed using artificial infection routes and artificial host–virus combinations. These may not reflect natural infections, especially for subtle phenotypes such as gene expression. Here, to explore the laboratory virus community and to better understand how natural virus infections induce changes in gene expression, we have analysed seven publicly available D. melanogaster transcriptomic sequencing datasets that were originally sequenced for projects unrelated to virus infection. We have found ten known viruses—including five that have not been experimentally isolated—but no previously unknown viruses. Our analysis of host gene expression revealed that numerous genes were differentially expressed in flies that were naturally infected with a virus. For example, flies infected with nora virus showed patterns of gene expression consistent with intestinal vacuolization and possible host repair via the upd3 JAK/STAT pathway. We also found marked sex differences in virus-induced differential gene expression. Our results show that natural virus infection in laboratory Drosophila does indeed induce detectable changes in gene expression, suggesting that this may form an important background condition for experimental studies in the laboratory.
Full article
(This article belongs to the Section Insect Viruses)
►▼
Show Figures
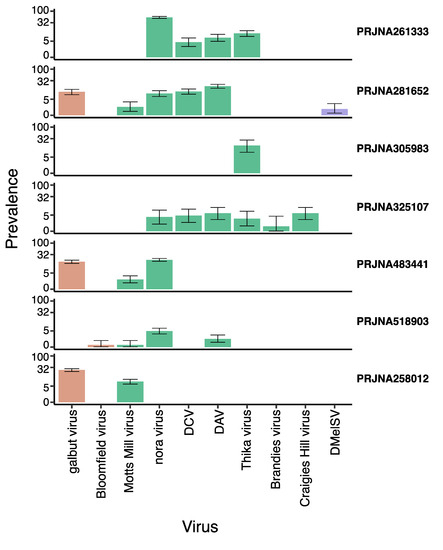
Figure 1
Open AccessArticle
Clinical Characterization of Respiratory Syncytial Virus Infection in Adults: A Neglected Disease?
by
, , , , , , , and
Viruses 2023, 15(9), 1848; https://doi.org/10.3390/v15091848 - 31 Aug 2023
Abstract
Lower respiratory tract infections (LRIs) are a significant cause of disability-adjusted life-years (DALYs) across all age groups, especially in children under 9 years of age, and adults over 75. The main causative agents are viruses, such as influenza and respiratory syncytial virus (RSV).
[...] Read more.
Lower respiratory tract infections (LRIs) are a significant cause of disability-adjusted life-years (DALYs) across all age groups, especially in children under 9 years of age, and adults over 75. The main causative agents are viruses, such as influenza and respiratory syncytial virus (RSV). Viral LRIs in adults have historically received less attention. This study investigated the incidence of RSV and influenza in adult patients admitted to a referral hospital, as well as the clinical profile of these infections. Molecular testing was conducted on nasopharyngeal samples taken from a respiratory surveillance cohort comprising adult (15–59 years) and elderly (60+ years) hospitalized patients who tested negative for SARS-CoV-2, to determine the prevalence for influenza and RSV. Influenza was found to be less frequent among the elderly. The main symptoms of RSV infections were cough, fever, dyspnea, malaise, and respiratory distress, while headache, nasal congestion, a sore throat, and myalgia were most frequent in influenza. Elderly patients with RSV were not found to have more severe illness than adults under age 60, underscoring the importance of providing the same care to adults with this viral infection.
Full article
(This article belongs to the Section Human Virology and Viral Diseases)
►▼
Show Figures
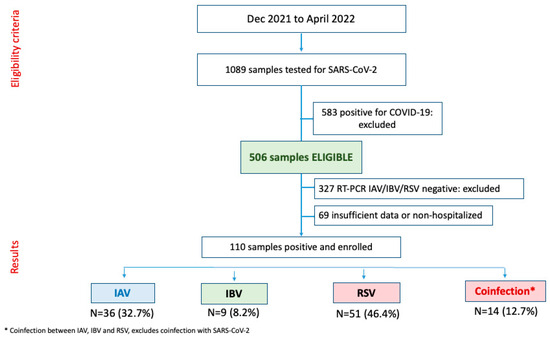
Figure 1
Open AccessReview
Feline Infectious Peritonitis: European Advisory Board on Cat Diseases Guidelines
by
, , , , , , , , , , , , , and
Viruses 2023, 15(9), 1847; https://doi.org/10.3390/v15091847 - 31 Aug 2023
Abstract
Feline coronavirus (FCoV) is a ubiquitous RNA virus of cats, which is transmitted faeco-orally. In these guidelines, the European Advisory Board on Cat Diseases (ABCD) presents a comprehensive review of feline infectious peritonitis (FIP). FCoV is primarily an enteric virus and most infections
[...] Read more.
Feline coronavirus (FCoV) is a ubiquitous RNA virus of cats, which is transmitted faeco-orally. In these guidelines, the European Advisory Board on Cat Diseases (ABCD) presents a comprehensive review of feline infectious peritonitis (FIP). FCoV is primarily an enteric virus and most infections do not cause clinical signs, or result in only enteritis, but a small proportion of FCoV-infected cats develop FIP. The pathology in FIP comprises a perivascular phlebitis that can affect any organ. Cats under two years old are most frequently affected by FIP. Most cats present with fever, anorexia, and weight loss; many have effusions, and some have ocular and/or neurological signs. Making a diagnosis is complex and ABCD FIP Diagnostic Approach Tools are available to aid veterinarians. Sampling an effusion, when present, for cytology, biochemistry, and FCoV RNA or FCoV antigen detection is very useful diagnostically. In the absence of an effusion, fine-needle aspirates from affected organs for cytology and FCoV RNA or FCoV antigen detection are helpful. Definitive diagnosis usually requires histopathology with FCoV antigen detection. Antiviral treatments now enable recovery in many cases from this previously fatal disease; nucleoside analogues (e.g., oral GS-441524) are very effective, although they are not available in all countries.
Full article
(This article belongs to the Special Issue Viral Infections in Companion Animals: Volume 2)
►▼
Show Figures
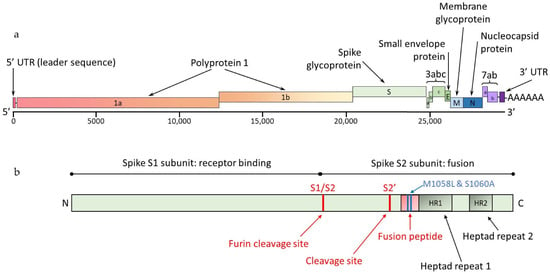
Figure 1

Journal Menu
► ▼ Journal Menu-
- Viruses Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Conferences
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Brain Sciences, Clinics and Practice, COVID, Life, Vaccines, Viruses
Multifaceted Efforts from Basic Research to Clinical Practice in Controlling COVID-19 Disease
Topic Editors: Yih-Horng Shiao, Rashi OjhaDeadline: 30 September 2023
Topic in
Agriculture, Agronomy, Crops, Plants, Viruses
Plant Virus
Topic Editors: Munir Mawassi, Sergey MorozovDeadline: 31 October 2023
Topic in
Biomolecules, BioTech, Marine Drugs, Microorganisms, Viruses
Marine Viruses
Topic Editors: Min Jin, Xiaobo Zhang, Rui Zhang, Claire GeslinDeadline: 30 November 2023
Topic in
Applied Sciences, Energies, Materials, Processes, Resources, Viruses
COVID-19 Impact on the Environment and Sustainable Development
Topic Editors: Elza Bontempi, Marco Race, Alessandra Zanoletti, Mario CocciaDeadline: 31 December 2023

Conferences
Special Issues
Special Issue in
Viruses
DNA Vaccines 2022
Guest Editor: Deborah H. FullerDeadline: 1 September 2023
Special Issue in
Viruses
Hepatitis B Virus: New Breakthroughs to Conquer an Ancient Disease
Guest Editors: Mark W. Douglas, Thomas TuDeadline: 15 September 2023
Special Issue in
Viruses
Human Paramyxoviruses
Guest Editors: Patrice Guillon, Larissa DirrDeadline: 30 September 2023
Special Issue in
Viruses
Diagnostics, Antiviral Therapy and Vaccination against Orthopoxvirus Infections
Guest Editors: Sergei N. Shchelkunov, Larisa N. ShishkinaDeadline: 15 October 2023
Topical Collections
Topical Collection in
Viruses
Coronaviruses
Collection Editors: Luis Martinez-Sobrido, Fernando Almazan Toral